Development of selective inhibitors for plant subtilisin-like serine proteases
SUPERVISOR: Lukas MACH
Background.
Several studies have demonstrated that plants are suitable hosts for the production of protein therapeutics (Sack et al., 2015). Unfortunately, the production of recombinant proteins in plants is often associated with unwanted proteolysis (Mandal et al., 2016).
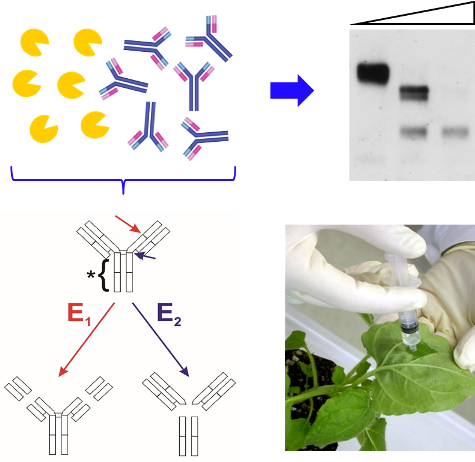
For recombinant monoclonal antibodies, the accumulation of substantial amounts of Fab- and Fc-like degradation products has been observed in several plant species including Nicotiana benthamiana (Hehle et al., 2015). Evidence has been obtained that in planta fragmentation of antibodies involves the action of serine proteases (Niemer et al., 2014). However, recent efforts to prevent proteolysis by co-expression of general serine protease inhibitors had only limited success (Grosse-Holz et al., 2018).
Aims and methods.
This project aims at the isolation and detailed characterization of high-affinity inhibitors for N. benthamiana subtilisin-like serine proteases capable of cleaving recombinant proteins such as monoclonal antibodies. Since recombinant protein degradation occurs largely in the apoplast (Castilho et al., 2014), we have recently purified the subtilisin-like serine proteases present in this compartment by capture with a selective activity-based probe. After identification by mass spectrometry and cloning of the corresponding cDNAs, the most abundant enzymes were ectopically expressed in N. benthamiana and purified exploiting their exogenously added affinity tags. Some of the recombinant proteases were found to cleave the anti-HIV antibodies 2F5 and PG9 at the same positions as unfractionated apoplastic fluid, suggesting that selective inhibition of the respective enzymes could increase the stability of recombinant proteins in planta.
A recent study has reported the occurrence of a tight-binding endogenous inhibitor of plant subtilisin-like serine proteases, which displays a much higher affinity for these enzymes than other serine protease inhibitors (Hohl et al., 2017). The first goal of this project is therefore the cloning of the N. benthamiana orthologue of this subtilisin propeptide-like inhibitor (NbSPI), which will then be functionally expressed in Escherichia coli. Purified NbSPI will be tested for its inhibitory capacity towards all recombinant N. benthamiana subtilisin-like serine proteases available in our laboratory. Inhibition kinetics will be determined by enzymatic assays based on the cleavage of specific internally quenched fluorescence substrates. Biophysical interaction studies will be performed by microscale thermophoresis. Molecular modelling based on the closely related cucumisin-propeptide structure (Sotokawauchi et al., 2017) will be used to identify NbSPI residues suitable for replacement. Efforts will then be undertaken to improve the affinity and selectivity of NbSPI for selected target proteases by saturation mutagenesis as well as by directed evolution based on a yeast surface display library approach. Binders will be isolated by fluorescence-activated cell sorting after incubation with catalytically inactive labelled variants of the respective bait enzymes (Cohen et al., 2018).
Finally, wild-type NbSPI and promising mutants will be assessed in co-expression experiments with the proteolysis-sensitive antibodies 2F5 and PG9 for their potential to down-regulate apoplastic serine protease activities and improve the yield and integrity of recombinant proteins. Antibody quality will be determined by mass spectrometry, differential scanning calorimetry and real-time antigen-binding assays based on surface plasmon resonance.
Collaborations within this thesis will include HALTRICH (saturation mutagenesis), OBINGER (directed evolution), OOSTENBRINK (molecular modelling) and STEINKELLNER (expression of recombinant proteins in plants). International collaborations include SCHALLER (biochemical and biophysical studies).
Castilho, A., Windwarder, M., Gattinger, P., Mach, L., Strasser, R., Altmann, F., Steinkellner, H. (2014) Proteolytic and N-glycan processing of human alpha1-antitrypsin expressed in Nicotiana benthamiana. Plant Physiol. 166, 1839-1851. doi: 10.1104/pp.114.250720
Cohen, I., Naftaly, S., Ben-Zeev, E., Hockla, A., Radisky, E.S., Papo, N. (2018) Pre-equilibrium competitive library screening for tuning inhibitor association rate and specificity toward serine proteases. Biochem. J. 475, 1335-1352. doi: 10.1042/BCJ20180070
Grosse-Holz, F., Madeira, L., Zahid, M.A., Songer, M., Kourelis, J., Fesenko, M., Ninck, S., Kaschani, F., Kaiser, M., van der Hoorn, R.A. (2018) Three unrelated protease inhibitors enhance accumulation of pharmaceutical recombinant proteins in N. benthamiana. Plant Biotechnol. J., doi: 10.1111/pbi.12916
Hehle, V.K., Lombardi, R., van Dolleweerd, C.J., Paul, M.J., Di Micco, P., Morea, V., Benvenuto, E., Donini, M., Ma, J.K. (2015) Site-specific proteolytic degradation of IgG monoclonal antibodies expressed in tobacco plants. Plant Biotechnol. J. 13, 235-245. doi: 10.1111/pbi.12266
Hohl, M., Stintzi, A., Schaller, A. (2017) A novel subtilase inhibitor in plants shows structural and functional similarities to protease propeptides. J. Biol. Chem. 292, 6389-6401. doi: 10.1074/jbc.M117.775445
Mandal, M.K., Ahvari, H., Schillberg, S., Schiermeyer, A. (2016) Tackling unwanted proteolysis in plant production hosts used for molecular farming. Front. Plant Sci. 7, 267. doi: 10.3389/fpls.2016.00267
Niemer, M., Mehofer, U., Torres Acosta, J.A., Verdianz, M., Henkel, T., Loos, A., Strasser, R., Maresch, D., Rademacher, T., Steinkellner, H., Mach, L. (2014) The human anti-HIV antibodies 2F5, 2G12, and PG9 differ in their susceptibility to proteolytic degradation: down-regulation of endogenous serine and cysteine proteinase activities could improve antibody production in plant-based expression platforms. Biotechnol. J. 9, 493-500. doi: 10.1002/biot.201300207
Sack, M., Hofbauer, A., Fischer, R., Stöger, E. (2015) The increasing value of plant-made proteins. Curr. Opin. Biotechnol. 32, 163-170. doi: 10.1016/j.copbio.2014.12.008
Sotokawauchi, A., Kato-Murayama, M., Murayama, K., Hosaka, T., Maeda, I., Onjo, M., Ohsawa, N., Kato, D.I., Arima, K., Shirouzu, M. (2017) Structural basis of cucumisin protease activity regulation by its propeptide. J. Biochem. 161, 45-53. doi: 10.1093/ jb/mvw053
