Allosteric molecular recognition to improve tumor specificity
PRINCIPAL INVESTIGATOR: Christian OBINGER
Background.
Adoptive transfer of T cells genetically modified to express chimeric antigen receptors (CARs) is one of the most promising strategies in the field of cancer immunotherapy. Despite impressive clinical responses with CAR T cells there are several challenges. For example, the treatment of solid tumors is currently limited by the lack of truly tumor-specific antigens. That is, virtually all currently targeted tumor antigens are also expressed on healthy tissues, which ultimately results in so-called on-target/off-tumor toxicity. One of the clinically best validated tumor-associated antigens is the epidermal growth factor receptor (EGFR), which is frequently mutated and/or up-regulated on various types of cancer. Again, however, virtually all EGFR-directed monoclonal antibodies (mAbs) are not specific for EGFR-molecules expressed on tumor cells.
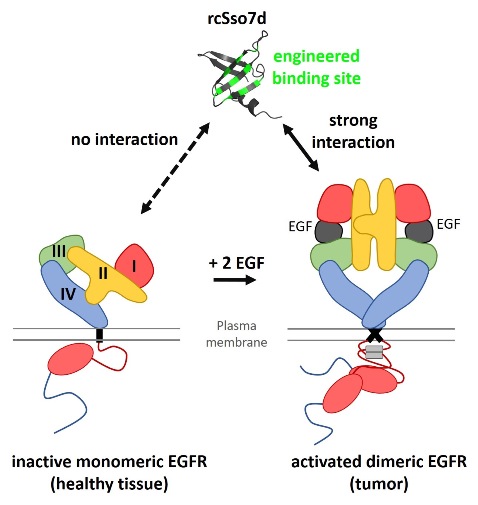
EGFR is a receptor tyrosine kinase, which undergoes a major conformational change upon ligand binding. Its extracellular domain thereby switches from a tethered to an extended, ligand-bound structure, which ultimately results in dimerization and activation of intracellular kinase function (Kovacs et al., 2015). Many cancers activate their EGFR molecules by autocrine signaling (Gazdar and Minna, 2008). Importantly, it is this activated state of EGFR – and not just its presence – which differentiates those tumor cells from normal, non-cancerous cells. Therefore, we hypothesize, that engineering protein-based binders for allosterically recognizing the activated, ligand-bound conformation of EGFR will improve the tumor specificity of EGFR-targeting approaches, such as EGFR-directed CAR T cells.
Aims and methods.
The extracellular domain of EGFR will be expressed as an IgG1-Fc fusion protein in HEK293T cells (cooperation with BORTH and MACH), as previously shown by our group (Traxlmayr et al., 2016). The purified EGFR-Fc protein will be used as the antigen for subsequent yeast display selections. More specifically, the binder scaffold rcSso7d will be engineered for binding to EGFR-Fc only in the presence, but not in the absence of the ligand EGF. We have already successfully engineered rcSso7d for EGFR-binding, demonstrating that the selection process using yeast display and EGFR-Fc as the antigen is feasible (Traxlmayr et al., 2016). The important modification in this project is the addition of EGFR-ligands during the yeast display experiment in order to direct the selection pressure towards recognition of ligand-bound (i.e. activated) EGFR. It is worth noting that the yeast display method has already been successfully applied in our laboratory (Traxlmayr et al., 2012 and 2013).
After yeast display selections, enriched rcSso7d-based EGFR-binders will be expressed solubly in E. coli and characterized in detail for their ability to bind to EGFR and – importantly – to differentiate between activated, ligand-bound EGFR and inactive EGFR. For those binding analyses surface plasmon resonance (SPR) will be applied in the absence and presence of EGFR-ligands. Moreover, the interaction between EGFR and the engineered binders will also be tested on the surface of human cells by flow cytometry, again in the presence and absence of EGFR-ligands. Furthermore, engineered binders will be analyzed for their tendency to aggregate and for their thermal stability by SEC and DSC, respectively. In order to investigate the molecular mechanism behind the specificity for ligand-bound EGFR, the complex structure of the most promising binder with EGFR-Fc will be solved by X-ray crystallography (cooperation with DJINOVIC-CARUGO). Structural studies will be complemented by MD simulations (cooperation with OOSTENBRINK). Finally, the engineered binders will be incorporated into CARs (cooperation with LEHNER). Primary human T cells transfected with the respective CAR constructs will be tested for their ability to specifically kill tumor cells expressing activated EGFR vs. non-cancerous cells that also express EGFR, but in an inactive state.
It is the clear goal of this project to investigate what degree of tumor specificity can be achieved by specifically engineering binders for allosteric recognition of the ligand-bound conformation of EGFR.
Collaborations within this thesis will include BORTH and MACH (mammalian cell factories) and OOSTENBRINK (MD simulations). National collaborations will include LEHNER (CAR T cells) and DJINOVIC-CARUGO (X-ray crystallography).
Gazdar, A.F. and Minna J.D. (2008) Deregulated EGFR Signaling during Lung Cancer Progression: Mutations, Amplicons, and Autocrine Loops. Cancer Prev. Res. 1, 156-160. doi: 10.1158/1940-6207.CAPR-08-0080
Kovacs, E., Zorn, J.A., Huang, Y., Barros, T., Kuriyan, J. (2015) A Structural Perspective on the Regulation of the Epidermal Growth Factor Receptor. Annu. Rev. Biochem. 84, 739-764. doi: 10.1146/annurev-biochem-060614-034402
Traxlmayr, M.W., Faissner, M., Stadlmayr, G., Hasenhindl, C., Antes, B., Rüker, F., Obinger, C. (2012) Directed evolution of stabilized IgG1-Fc scaffolds by application of strong heat shock to libraries displayed on yeast. Biochim. Biophys. Acta 1824, 542-549. doi: 10.1016/j.bbapap.2012.01.006
Traxlmayr, M.W., Lobner, E., Antes, B., Kainer, M., Wiederkum, S., Hasenhindl, C., Stadlmayr, G., Rüker, F., Woisetschläger, M., Moulder, K., Obinger, C. (2013) Directed evolution of Her2/neu-binding IgG1-Fc for improved stability and resistance to aggregation by using yeast surface display. Prot. Eng. Des. Sel. 26, 255-265. doi: 10.1093/protein/gzs102
Traxlmayr, M.W., Kiefer, J.D., Srinivas, R.R., Lobner, E., Tisdale, A.W., Mehta, N.K., Yang, N.J., Tidor, B., Wittrup, K.D. (2016) Strong Enrichment of Aromatic Residues in Binding Sites from a Charge-neutralized Hyperthermostable Sso7d Scaffold Library. J. Biol. Chem. 291, 22496-22508. doi: 10.1074/jbc.M116.741314
