Characterization and engineering of microbial methanol metabolism
SUPERVISOR: DIETHARD MATTANOVICH
Background.
A number of microorganisms have evolved pathways to utilize one−carbon substrates like methanol as sources for energy and carbon (Anthony, 1991). Among them yeasts and some methylotrophic bacteria share the common pathway that methanolis first oxidized to formaldehyde and then either assimilated via the pentose phosphate pathway, or further oxidized to CO2 for energy production. Assimilation further uses intermediates of gluconeogenesis and the pentose phosphate pathway to create the sugars needed for cellular growth (Schrader et al., 2009).
The major difference between yeasts and bacteria is the efficiency of the first step, oxidation of methanol. Yeasts use alcohol oxidases (EC 1.1.3.13, prosthetic group: FAD) for this step, which transfer the electrons to oxygen thereby producing hydrogen peroxide (Geissler and Hemmerich, 1981), which burdens the cells and must be degraded by other enzymes like catalase and peroxidases. By contrast, most methylotrophic bacteria use NAD(P)+ dependent alcohol dehydrogenases (EC 1.1.1.2, cofactor: Zn2+) which transfer electrons to NAD+ or NADP+, respectively (Krog et al., 2013). This has major physiological consequences: alcohol oxidases cannot make use of the energy and reductive power that is liberated by oxidation of methanol. Consequently large amounts of metabolic heat are liberated, and the biomass yield on methanol is only about half of the theoretical yield (Mattanovich et al., 2014). Methanol metabolism is therefore much more efficient in bacteria than in yeasts.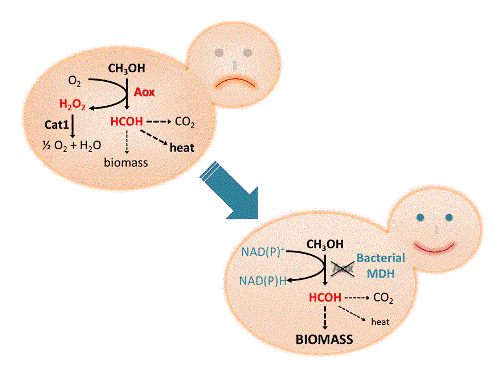
Aims and methods.
The objectives of this project are to study the differences of bacterial and yeast methanol metabolism by (1) characterizing respective methanol−oxidizing enzymes, and (2) unifying both pathways in one system, the yeast P. pastoris. The metabolic impact of the additional gain of energy and reducing power by methanol dehydrogenases will be characterized, and the difference in metabolic heat production and oxygen consumption will be quantified.
In a first step suitable bacterial methanol dehydrogenases will be selected and bacterial enzyme preparations will be screened in vitro for co−factor utilization (NAD+ or NADP+), specific activity, pH and temperature optimum, as well as conformational stability (OBINGER) in order to identifiy the best suited enzyme to work in yeast. Simultaneously, methanol metabolism, cell growth and protein production will be simulated in silico by metabolic modelling to identify which co−factor should work best and which fluxes through methanol oxidation appear feasible (ZANGHELLINI). An AOX double knock out (mut−) of P. pastoris will be created (GASSER). The selected bacterial methanol dehydrogenases will be cloned under different P. pastoris promoters (regulated and constitutive) of different strength (Prielhofer et al., 2013; Delic et al., 2013), and with different intracellular targeting signals (to cytosol and peroxisomes). After screening for growth on methanol and high biomass productivity metabolic fluxes through methanol assimilation and central carbon metabolic pathways will be measured as well as the status of catabolic and anabolic reduction charges (NAD/H and NADP/H; Ortmayr et al., 2014). The impact on amino acid synthesis and energy production will reflect changes in assimilatory and dissimilatory processes due to more efficient substrate utilization.
Anthony, C. (1991) Assimilation of carbon by methylotrophs. Biotechnology 18, 79–109
Krog, A., Heggeset, T.M., Mueller, J.E., Kupper, C.E., Schneider, O., Vorholt, J.A., Ellingsen, T.E., Brautaset, T. (2013) Methylotrophic Bacillus methanolicus encodes two chromosomal and one plasmid born NAD+ dependent methanol dehydrogenase paralogs with different catalytic and biochemical properties. PLoS One 8:e59188
Mattanovich, D., Jungo, C., Wenger, J., Dabros, M., Maurer, M. (2014) Yeast Suspension Culture. In: Hans-Peter Meyer, Diego Schmidhalter (Eds) Industrial Scale Suspension Culture of Living Cells, Wiley-Blackwell
Schrader, J., Schilling, M., Holtmann, D., Sell, D., Filho, M.V. (2009) Methanol-based industrial biotechnology: current status and future perspectives of methylotrophic bacteria. Trends Biotechnol. 27, 107–115
Ortmayr, K., Nocon, J., Gasser, B., Mattanovich, D., Hann, S., Koellensperger, G. (2014) Sample preparation workflow for the LC-MS based analysis of NADP+ and NADPH in yeast. J. Sep. Sci., doi: 10.1002/jssc.201400290
Geissler, J., Hemmerich, P. (1981) Yeast methanol oxidases: an unusual type of flavoprotein. FEBS Lett. 126, 152-156
