Establishment of a hyper-secreting eukaryotic protein production system as alternative cell factory for biotherapeutics
SUPERVISOR: NICOLE BORTH
Background.
Recombinant therapeutic proteins are currently produced primarily in mammalian cell factories, specifically in Chinese Hamster Ovary cells. These cells, while highly flexible and able to produce safe and high quality proteins, also have a number of disadvantages. These include high genomic and phenotypic variation, instability of production, secretory bottlenecks and interclonal variation. These problems are not likely to be overcome by switching to any other mammalian cell system, as inherently, due to their large genomes and complex regulatory networks, these have the same issues of genomic instability and phenotypic variation. An alternative system would have to fulfil the following prerequisites: a small, diploid genome (<1Gb); absence of a cell wall; simple, chemically defined and cost efficient growth media; fast growth rate (less than 15h per division); efficient and highly productive protein synthesis and secretion; eukaryotic organism (as ER and Golgi are required for proper assembly and post-translational modification of most secretory proteins). Other requirements, such as human like glycosylation, or existing selection systems for genetic engineering, are less of an issue in times of efficient genome editing, as these can be engineered into the cell at a later stage.
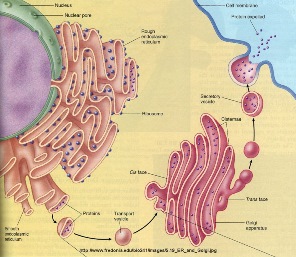
Aims and methods.
The aim of this project is to evaluate different organisms and cell lines that might be used to replace current mammalian cell factories. We have recently developed a protocol for transient transfection of mRNA, which is dosable and enables reaching industrially relevant high production rates for short periods of time (24-48h). This protocol will be adapted to test for the biomass specific maximum protein secretion rate of a given organism or cell line in a standardised and quantitative way from a defined number of mRNA copies. The initial task will be to determine the maximum protein secretion rate of existing cell lines and cell factories as a quantitative background measure to evaluate novel candidates. In subsequent work, we will look into the mechanism underlying the higher yield of secreted protein in the best candidates: be it higher translation rates, better assembly or more efficient vesicular transport.
Subsequently, novel organisms and cell types will be evaluated for their potential as cell factories. Organisms of interest include insects, protists and small marine organisms. We have already identified a number of organisms or cell lines which could fulfil the criteria defined above (Dictyostelium (as a protist example), several existing insect cell lines such as S2 (Drosophila) or Tn5 (Trichoplusia ni) and Labyrinthula as an example for marine organisms.
Selected candidates will be screened for natural protein secretion by analysing the amount of total protein in the supernatant of cultures and relating it to cell number and then tested for maximum achievable transient protein secretion as described above. Promising candidates will be further evaluated for growth, final cell density, available culture media, genome size, available selection systems and DNA transfectability.
Depending on progress, the final outcome will be a list of candidate cell lines or organisms with optimised media, defined protocols for DNA transfection and an optimised selection system (if necessary engineered into the cells). Scientifically, the main output will be a ranking of cell lines and organisms according to maximum possible protein secretion rates that will compare tested candidates to available cell factories on a per biomass basis.
