Quantitative metabolic flux analysis in CHO cells
SUPERVISOR: NICOLE BORTH
Background.
Chinese hamster ovary cells are the primary host organisms for the production of therapeutic proteins, mainly because of their ability to perform human-like glycosylation, ease of adaptation to serum-free growth in suspension, and safety. The increasing demand for biotherapeutics requires the development of more efficient producer strains and bioprocess strategies (1). Research so far has been mostly focused on optimizing the bioprocess. With the availability of the genome sequence of CHO as well as precise genome-editing tools such as CRISPR/Cas9, metabolism has now become an attractive target to further optimize product titres and protein glycosylation quality. Metabolic flux analysis has proved to be an efficient strategy to gain deeper knowledge about cell metabolism. It allows for the identification of possible metabolic engineering targets, the prediction of cellular behaviour during the bioprocess, the optimization of media formulations and the rational design of feed and control strategies (2).
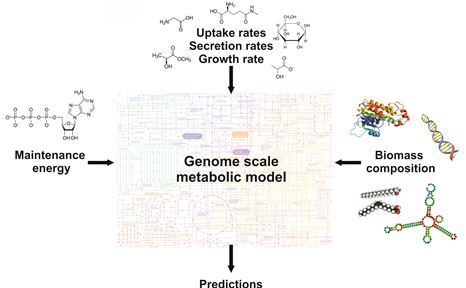
Aims and methods.
The aims of this research include the understanding of metabolic changes in CHO strains adapted to growth without glutamine and the determination of the biomass compositions and maintenance energies of several CHO strains.
Recently, non-producer and producer CHO cell lines were adapted to growth without glutamine (3, 4), in order to reduce the negative effects of ammonia produced by glutaminolysis (5, 6). Fermentations with various CHO strains under different conditions will be run, gathering data such as growth, pH, concentration of extracellular metabolites and product titre (in case of producer strains). The obtained data will be used as input for flux balance analysis (FBA) with a recently created CHO specific genome-scale metabolic model. Metabolic differences between the wildtype strains and the strains adapted to growth without glutamine, for both the producer and the non-producer strains, will be investigated.
The current constraints of the project come from lack of information on the precise biomass composition of CHO cells and its variation, and, in addition, from missing data on maintenance energy. Both of these data sets are required for correct calculation of flux balances and have been shown to make precise predictions impossible if incorrect or lacking (7, 8, 9). The project will therefore address this need by generating the required data sets and will then use them on a model system. Biomass composition of CHO cells, its variability among different CHO strains and changes during various stages of cultivation will be determined. Maintenance energy coefficient will be calculated by performing continuous fermentations with various dilution rates and measuring the concentration of ATP (and other NTPs).
1. Kim, J. Y., Kim, Y. G., & Lee, G. M. (2012). CHO cells in biotechnology for production of recombinant proteins: current state and further potential. Appl Microbiol Biotechnol, 93(3), 917-930.
2. Zupke, C., & Stephanopoulos, G. (1995). Intracellular flux analysis in hybridomas using mass balances and in vitro (13)C nmr. Biotechnol Bioeng, 45(4), 292-303.
3. Bort, J. A., Stern, B., & Borth, N. (2010). CHO-K1 host cells adapted to growth in glutamine-free medium by FACS-assisted evolution. Biotechnol J, 5(10), 1090-1097.
4. Taschwer, M., Hackl, M., Hernández Bort, J. A., Leitner, C., Kumar, N., Puc, U., et al. (2012). Growth, productivity and protein glycosylation in a CHO EpoFc producer cell line adapted to glutamine-free growth. J Biotechnol, 157(2), 295-303.
5. Hansen, H. A., & Emborg, C. (1994). Influence of ammonium on growth, metabolism, and productivity of a continuous suspension Chinese hamster ovary cell culture. Biotechnol Prog, 10(1), 121-124.
6.Yang, M., & Butler, M. (2000). Effects of ammonia on CHO cell growth, erythropoietin production, and glycosylation. Biotechnol Bioeng, 68(4), 370-380.
7.Pramanik, J., & Keasling, J. D. (1998). Effect of Escherichia coli biomass composition on central metabolic fluxes predicted by a stoichiometric model. Biotechnol Bioeng, 60(2), 230-238.
8.Selvarasu, S., Ho, Y. S., Chong, W. P., Wong, N. S., Yusufi, F. N., Lee, Y. Y., et al. (2012). Combined in silico modeling and metabolomics analysis to characterize fed-batch CHO cell culture. Biotechnol Bioeng, 109(6), 1415-1429.
9.Kliphuis, A. M., Klok, A. J., Martens, D. E., Lamers, P. P., Janssen, M., & Wijffels, R. H. (2012). Metabolic modeling of Chlamydomonas reinhardtii: energy requirements for photoautotrophic growth and maintenance. J Appl Phycol, 24(2), 253-266.
