Generation and presentation of membrane protein targets using influenza virus−like particles
SUPERVISOR: REINGARD GRABHERR
Background.
Membrane proteins play important roles in performing many cellular processes and thus, are one of the most important targets in cancer therapy. Hence, these proteins are often used as targets for antibody-based therapies. However, membrane proteins are notoriously difficult to produce and study, due to their hydrophobic nature and complex structures. In contrast, secreted proteins and virus-like particles are efficiently produced in insect cells and transported to the cell culture supernatant (Krammer et al., 2010a). Influenza virus-like particles have successfully been produced in baculovirus-based expression systems, displaying different types of hemagglutinin (Krammer et al., 2010b; Klausberger et al., 2014).
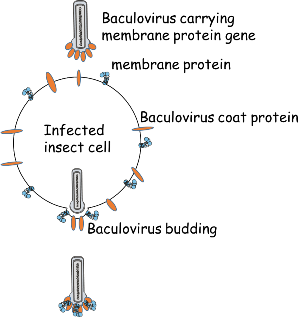
In the course of this work, we could show that for this purpose Trichoplusia ni cell lines are advantageous as compared to other insect cell lines (Wilde et al., 2014). In contrast to baculoviruses, influenza virus particles also form budded virions when the envelope protein hemagglutinin is absent and solely the matrix protein M1 is expressed. Thus, when baculoviruses contain the M1 gene in combination with a membrane protein gene of choice, budded virion particles with desired surface properties can be harvested from the cell culture supernatant of Trichoplusia ni cell lines.
Aims and methods.
The goal of this project is to use influenza virus-like particles as display scaffold for relevant biomarkers in such a manner that screening assays can be performed (e.g. against bacterial surface display libraries). We will focus on two target proteins: the epidermal growth factor receptor (EGFR) and epidermal growth factor receptor 2 (HER2). Mutations that lead to EGFR overexpression have been associated with a number of cancers, including lung cancer and glioblastoma multiforme, thus making EGFR an interesting target for cancer therapy (Kuan et al., 2001). As with breast cancer, patients with gastric and gastro-oesophageal junction cancers overexpress HER2 in 10−15% of cases (Smyth et al., 2005). Hence, patients with advanced HER2-positive tumors benefit from treatment with anti-HER2 antibodies (Bang et al., 2010).
Full-length EGFR and HER2 will be expressed in insect cells, and protein incorporated into the membrane of ted cells as well as budded virus-like particles, will be analysed in terms of avidity, structural properties and glycosylation. After purification (JUNGBAUER), these particles serve as screening targets for bacterial phage display libraries expressing antibody fragments. Screening experiments will be done in collaboration with John McCafferty, University of Cambridge, and BORTH. If required, glycomodulation will be performed in order to simulate human glycan patterns using the Sweetbac technology (Palmberger et al., 2012a, Palmberger et al., 2012b). Altered glycan structures will be analysed by mass spectrometry (ALTMANN).
Cragg, M. S., Walshe, C. A., Ivanov, A. O., Glennie, M. J. (2005) The biology of CD20 and its potential as a target for mAb therapy. Curr. Dir. Autoimmun. 8, 140-174
Grabherr, R., Ernst, W. (2002) Baculovirus major envelope protein gp64 and its application in eukaryotic surface display strategies. Recent Res. Dev. Biochem. 3, 445-457
Hulett, M., Hogarth, P. (1998) The second and third extracellular domains of FcγRI (CD64) confer the unique high affinity binding of IgG2a. Mol. Immunol. 35, 989–996
Kuan, C. T., Wikstrand, C. J., Bigner, D. D. (2001) EGF mutant receptor vIII as a molecular target in cancer therapy. Endocr. Relat. Cancer 8, 83–96
Lai, Y.-K., Lai, C.-W., Liu, H.-J., Hu, Y.-C. (2007) Avian influenza virus hemagglutinin display on baculovirus envelope: Cytoplasmic domain affects virus properties and vaccine potential. Mol. Ther. 15, 989-996
Lu, L., Yu, L., Kwang, J. (2007) Baculovirus surface-displayed hemagglutinin of H5N1 influenza virus sustains its authentic cleavage, hemagglutination activity, and antigenicity. Biochem. Biophys. Res. Commun. 358, 404-409
Mangor, J. T., Monsma, S. A., Johnson, M. C., Blissard, G. W. (2001). A GP64-null baculovirus pseudotyped with vesicular stomatitis virus G protein. J. Virol. 75, 2544-2556
Krammer, F., Grabherr, R. (2010a) Alternative influenza vaccines made by insect cells. Trends Mol. Med. 16, 313-320
Krammer, F., Palmberger, D., Tauer, C., Messner, P., Grabherr, R. (2010b) Trichoplusia ni cells (high-five) are highly efficient for the production of Influenza A virus-like particles; a comparison of two insect cell lines as production platforms for influenza vaccines. Mol. Biotechnol. 45, 226-234
Klausberger, M., Wilde, M., Palmberger, D., Hai, R., Albrecht, R. A., Margine, I., Hirsh, A., Grabherr, R., Krammer, F. (2013) One-shot vaccination with an insect cell-derived low-dose influenza A H7 virus-like particle preparation protects mice against challenge. Vaccine 32, 355-362
Palmberger, D., Klausberger, M., Berger, I., Grabherr, R. (2012a) MultiBac turns sweet. Bioengineered 4, 1-6
Palmberger, D., Wilson, I. B. H., Berger, I., Grabherr, R., Rendic, D. (2012b) SweetBac: A new approach for the production of mammalianised glycoproteins in insect cells. PloS One 7:e34226
Wilde, M., Klausberger, M., Palmberger, D., Ernst, W., Grabherr, R. (2014) Tnao38, High Five and Sf9 − Evaluation of host virus interactions in three different insect cell lines: baculovirus production and recombinant protein expression. Biotechnol. Lett. 36, 743-749
Smyth, E. C., Cunningham, D. (2012) Targeted therapy for gastric cancer. Curr. Treat. Options Oncol. 13, 377-389
Bang, Y. J., van Clutsem, F., Feyereislova, A. et al., (2010) ToGA Trial Investigations. Trastuzumab in combination with chemotherapy versus chemotherapy alone for treatment of HER2-positive advanced gastric or gastric-oesophageal junction cancer (ToGA): a phase 3, open-label, randomised controlled trial. Lancet 376, 687-697. Erratum in Lancet. 2010; 376, 1302
