Structure⁄function relationships in the electron transfer reactions of cellobiose dehydrogenase
PRINCIPAL INVESTIGATOR: DIETMAR HALTRICH
Background.
Cellobiose dehydrogenase (CDH) is an extracellular enzyme produced under cellulolytic culture conditions by a number of lignocellulose-degrading fungi from the phyla Basidiomycota and Ascomycota (Zamocky et al., 2006). CDH (CAZy family AA3_1) typically is a 80–100 kDa large extracellular flavocytochrome featuring a haem-b-containing cytochrome domain (CYT) connected by a ∼15-residue linker to a flavin-adenine dinucleotide (FAD)-binding dehydrogenase domain (DH), the latter belonging to the glucose-methanol-choline family of oxidoreductases. The DH domain oxidizes cellobiose (and other sugars) at the C1 position to cellobiono-1,5-lactone whereby the FAD cofactor is reduced. The ensuing step involves inter-domain electron transfer from reduced FAD to the haem b of CYT, presumably by two single electron-transfer events, followed by an electron transfer from CYT to external electron acceptors. Only recently, an interaction between CDH and lytic polysaccharide mono-oxygenases (LPMO) has been shown, with LPMOs serving as a possible acceptor for electrons from CDH (Sygmund et al., 2012).
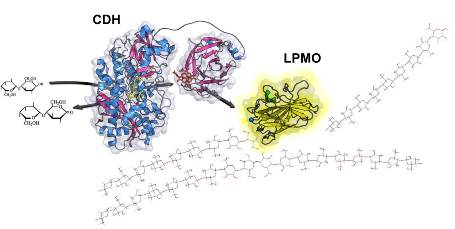
Fungal LPMOs (CAZy family AA9) are typically small (∼20 kDa), copper-containing enzymes that are ubiquitous in filamentous fungi. They have only recently been found to play a central role in oxidative degradation of cellulose, and to have potential in the conversion of plant biomass to commodity chemicals such as biofuels since they boost the decomposition of recalcitrant crystalline cellulose significantly (Horn et al., 2012). Crystal structures of several LPMOs reveal an absolutely conserved copper-containing active site coordinated by two His residues on a flat protein surface. LPMOs are proposed to bind and then cleave glycosidic bonds of cellulose, generating new chain ends on the cellulose surface that are subsequently processed by other (hydrolytic) cellulases. LPMOs require electrons for their oxidative reaction, with the in vivo source of these electrons still unknown. CDH is a very likely candidate for the provision of these electrons to LPMO (Phillips et al., 2011; Beeson et al., 2012), yet very little is known about the inter-domain electron transfer (DH to CYT domain), or the interaction and electron transfer between CDH and LPMO.
Our project partner Prof. Divne only recently solved the structures of intact CDH from two fungal sources (Myriococcum thermophilum and Neurospora crassa; manuscript submitted for publication) as well as LPMO from N. crassa; these enzymes being produced recombinantly in our group (Zamocky et al., 2008; Kittl et al., 20012; Sygmund et al., 2012). These CDH structures were obtained both in an open and closed form, the latter giving structural information about the interaction between the two domains and a possible electron transfer path between the two prosthetic groups. Knowledge about the electron pathway is furthermore of interest since CDH can transfer electrons directly via its CYT domain (without the application of a mediator) to electrodes, which is of interest for the application in biosensors and biofuel cells (Tasca et al., 2009).
Aims and methods.
This project aims at elucidating the electron transfer pathways from the flavin to the cytochrome domain in CDH as well as from the haem b in CDH to the copper in LPMO as well as amino acid residues involved in these pathways. Preliminary data obtained in our laboratory indicate that the inter-domain transfer (from the FAD to the haem) is the rate-limiting step in this overall process, so this process will be considered first. The interdomain electron transfer can be easily studied, since the electron acceptor cytochrome c is exclusively reduced at the CYT domain of CDH. Based on the now known structures of intact CDH both in the open and closed form we can rationally engineer possible electron pathways or identify factors that affect the binding / interaction of the two domains. An important input will be from OOSTENBRINK who will be using computational methods based on the new CDH structures available to study the domain interaction and electron transfer, using amongst others approaches that have been successfully applied to elucidate the electron pathway in the multidomain protein cytochrome P450 reductase (Suendermann and Oostenbrink, 2013). Amino acids identified in these modeling studies as important for domain interaction / electron transfer will be mutated (alanine scanning) to prove their importance, or replaced by selected amino acids that might improve these processes. Selected variants will be produced in Pichia pastoris and studied by biochemical, bioelectrochemical and biophysical methods including rapid kinetics (determination of rates for the electron transfer) as well as EPR (domain mobility and movement) together with OBINGER and GORTON.
In the second phase we will study the interaction and electron transfer from CDH, i.e., its CYT domain, to selected LPMOs, e.g., belonging to different classes (Class 1, 2, 3; Vu et al., 2014), which are characterised by distinctive active site surface features (LUDWIG). A starting point for these studies is N. crassa LPMO9F, for which our external project partner DIVNE deduced the structure, or other LPMOs with published structural data (Hemsworth et al., 2013). Based on the crystal structures of CDH and LPMO we proposed a direct electron transfer between haem b (or more precisely the propionate A) and the copper in LPMO. However, the copper is centered in a flat surface of LPMO, which is facing the surface of cellulose during catalysis and thus blocked. It will be of particular interest to search whether alternative electron pathways to the LPMO active site are feasible by using molecular modeling. Again, interesting variants (proving or disproving the involvement of certain amino acids in the proposed e-transfer pathway) will be produced in P. pastoris and studied together with EIJSINK.
Beeson, W.T., Phillips, C.M., Cate, J.H., Marletta, M.A. (2012) J. Am. Chem. Soc. 134, 890-892
Harreither, W., Sygmund, C., Augustin, M., Narciso, M., Rabinovich, M.L., Gorton, L., Haltrich, D., Ludwig, R. (2011) Catalytic properties and classification of cellobiose dehydrogenases from ascomycetes. Appl. Environ. Microbiol. 77, 1804–1815
Hemsworth, G.R., Davies, G.J., Walton, P.H. (2013) Curr. Opin. Struct. Biol. 23, 660–668
Horn, S.J., Vaaje-Kolstad, G., Westereng, B., Eijsink, V.G. (2012) Biotechnol. Biofuels 5, 45
Kittl, R., Kracher, D., Burgstaller, D., Haltrich, D., Ludwig, R. (2012) Biotechnol. Biofuels 5, 79
Phillips, C.M., Beeson, W.T., Cate, J.H., Marletta, M.A. (2011) ACS Chem. Biol. 6, 1399–1406
Suendermann A., Oostenbrink, C. (2013) Protein Sci. 22, 1183–1195
Sygmund, C., Kracher, D., Scheiblbrandner, S., Zahma, K., Felice, A.K.G., Harreither, W., Kittl, R., Ludwig, R. (2012) Appl. Environ. Microbiol. 78, 6161−6171
Tasca, F., Gorton, L., Harreither, W., Haltrich, D., Ludwig, R., Noell, G. (2009) Anal. Chem. 81, 2791-2798
Vu, V.V., Beeson, W.T., Phillips, C.M., Cate, J.H.D., Marletta, M.A. (2014) J. Am. Chem. Soc. 136, 562−565
Zamocky, M., Ludwig, R., Peterbauer, C., Hallberg, B.M., Divne, C., Nicholls, P., Haltrich, D. (2006) Curr. Protoc. Pept. Sci. 7, 255–280
Zamocky, M., Schuemann, C., Sygmund, C., O′Callaghan, J., Dobson, A., Ludwig, R., Haltrich, D., Peterbauer, C. (2008) Protein Expr. Purif. 59, 258-265
