Genomic and epigenomic analysis of a plant-based expression platform for recombinant glycoproteins
JOINT-SUPERVISORS: Heinz HIMMELBAUER, Lukas MACH
Background.
The tobacco-related plant species Nicotiana benthamiana is emerging as a promising host for the rapid, versatile and safe production of biopharmaceuticals. Recent advances in expression technologies allow the synthesis of monoclonal antibodies and other therapeutic proteins in N. benthamiana on an industrially relevant scale (Qiu et al., 2014). Furthermore, N. benthamiana lines have been established which permit the production of recombinant glycoproteins with customized N- and O-glycan structures and thus superior biological activities (Castilho et al., 2012; Strasser et al., 2014). However, a significant problem frequently encountered during the production of therapeutic proteins in Nicotiana species is their proteolytic degradation within the plants (Niemer et al., 2014). Although a draft genome sequence of the N. benthamiana line Nb-1 has been recently released (Bombarely et al., 2012), the spectrum of proteases present in this plant species is so far only superficially characterized (Goulet et al., 2012). There is also considerable lack of knowledge with respect to a number of enzymes involved in undesirable protein and glycan modifications such as the hydroxylation and subsequent glycosylation of proline residues (Castilho et al., 2012).
Aims and methods.
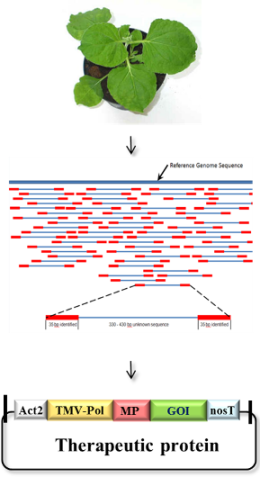
This project will focus on a detailed genomic and epigenomic analysis of a glycoengineered N. benthamiana line optimized for the production of recombinant proteins and its comparison with wild-type plants. For this, we will rely on our broad experience with the acquisition of next-generation sequencing (NGS) datasets and their utilization for the assembly and annotation of complex plant genomes (Dohm et al., 2014). NGS libraries will be generated from genomic DNA and mRNA isolated from leaves and other organs. Computational analysis of the data obtained with these sequencing libraries will then involve mapping of read-pairs against the Nb-1 draft genome, with the aim of assessing the genetic differences existing between Nb-1 and glycoengineered N. benthamiana, as well as its progenitor line. For once, this information can be used to reconstruct the genome sequence of glycoengineered N. benthamiana. Secondly, these studies will yield a comprehensive assessment of the genomic status of the currently most frequently used N. benthamiana production line and reveal any genetic or epigenetic alterations incurred during its generation. Special attention will be paid to the identification and transcriptional activity of genes encoding cysteine and serine proteinases, since degradation of recombinant proteins in planta is largely due to members of these protease families (Niemer et al., 2014). Particular emphasis will be also placed on the characterization of the N. benthamiana complement of glycosyltransferases, glycosidases and prolyl hydroxylases whose activities could potentially lead to the formation of non-human N- and O-glycan structures or other protein modifications on biotherapeutics (Castilho et al., 2012). Altogether, the outcomes of this project will provide a powerful resource for the further development of this attractive new expression platform.
Collaborations within this thesis will include ALTMANN (prolyl hydroxylases), MACH (proteases), STEINKELLNER (glycosidases), STRASSER (glycosyltransferases), and DOHM (genomics and transcriptomics).
Bombarely A, Rosli HG, Vrebalov J, Moffett P, Mueller LA, Martin GB (2012) A draft genome sequence of Nicotiana benthamiana to enhance molecular plant-microbe biology research. Mol. Plant Microbe Interact. 25, 1523-30
Castilho A, Neumann L, Daskalova S, Mason HS, Steinkellner H, Altmann F, Strasser R (2012) Engineering of sialylated mucin-type O-glycosylation in plants. J. Biol. Chem. 287, 36518-26
Dohm JC, Minoche AE, Holtgraewe D, Capella-Gutierrez S, Zakrzewski F, Tafer H, Rupp O, Soerensen TR, Stracke R, Reinhardt R, Goesmann A, Kraft T, Schulz B, Stadler PF, Schmidt T, Gabaldon T, Lehrach H, Weisshaar B, Himmelbauer H (2014) The genome of the recently domesticated crop plant sugar beet (Beta vulgaris). Nature 505, 546-9
Goulet C, Khalf M, Sainsbury F, D'Aoust MA, Michaud D (2012) A protease activity-depleted environment for heterologous proteins migrating towards the leaf cell apoplast. Plant Biotechnol. J. 10, 83-94
Niemer M, Mehofer U, Torres Acosta JA, Verdianz M, Henkel T, Loos A, Strasser R, Maresch D, Rademacher T, Steinkellner H, Mach L (2014) The human anti-HIV antibodies 2F5, 2G12, and PG9 differ in their susceptibility to proteolytic degradation: down-regulation of endogenous serine and cysteine proteinase activities could improve antibody production in plant-based expression platforms. Biotechnol. J. 9, 493-500
Qiu X, Wong G, Audet J, Bello A, Fernando L, Alimonti JB, Fausther-Bovendo H, Wei H, Aviles J, Hiatt E, Johnson A, Morton J, Swope K, Bohorov O, Bohorova N, Goodman C, Kim D, Pauly MH, Velasco J, Pettitt J, Olinger GG, Whaley K, Xu B, Strong JE, Zeitlin L, Kobinger GP (2014) Reversion of advanced Ebola virus disease in nonhuman primates with ZMapp. Nature 514, 47-53
Strasser R, Altmann F, Steinkellner H (2014) Controlled glycosylation of plant-produced recombinant proteins. Curr. Opin. Biotechnol. 30, 95-100
