In silico guided humanization of antibodies
SUPERVISOR: RENATE KUNERT
Background.
Monoclonal antibodies are still and increasingly in the focus of therapeutic application since inflammatory and immunological diseases as well as tumors have been successfully treated with such sophisticated proteins or chimera thereof (fusion proteins). Often, the antibodies are developed in scientific research institutes and small startup companies by immunization of lab animals. After promising in vitro and limited clinical trials the rodent derived antibodies need to be humanized to circumvent human anti-mouse immunogenicity after a couple of treatments. Our group has a long history in expression and investigation of various subtypes of mAbs and also focuses on humanization strategies. In close cooperation with the institute of molecular modelling and simulation at BOKU, we developed a computational approach to prioritize humanization candidates and avoid rather time consuming and lab intensive expression steps.
Aims and methods. 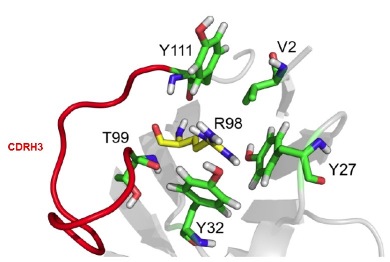
In this project, we will use the murine mAb Ab2/3H6 as a model antibody. In 2008, Bryson et al. investigated the interaction between the anti-idiotype Ab2/3H6 and the original anti HIV-1 mAb 2F5 (both developed at the Department of Biotechnology) by X-ray crystallography (Bryson et al., 2008) and determined characteristic residues responsible for the interaction. In parallel, we defined the most important amino acid at the CDR3 loop of Ab2/3H6 HC necessary for binding to 2F5. Computer models demonstrated the structural background (Gach et al., 2008; De Ruiter et al. 2011)
The humanization of Ab2/3H6 was initiated with different approaches, such as CDR-grafting, resurfacing and superhumanization (Mader and Kunert, 2010). The superhumanized variant was found to be inactive, but with the aid of computer simulations, a single mutation was identified that could restore some binding strength. The data gained from this study encouraged us to define the following work packages for the current PhD project:
Further investigation of the role of R98(H) for the proper orientation of CDR3. The previous work showed a remarkable influence of the residue 98 in the heavy chain. The T98R backmutation restored binding affinity of the variant. In the mouse structure, R98(H) is placed in the middle of three Tyr residues. Alternative variants may be proposed to investigate the highly relevant role of R98(H), such as a K98 variant to maintain stability, various mutations of the Tyr-cluster to disrupt stability, or the introduction of negatively charged residues at these positions to increase stability.
Improving binding of the superhumanized T98R mutant towards wild-type affinity.
While the single backmutation T98R shows a remarkable restoration of binding affinity from the inactive superhumanized variant, the RA02 mutant shows a binding affinity of - 39 kJ/mol, as compared to -51 kJ/mol for the murine wild-type. This indicates that position 98 is a relevant, but not the only determinant for affinity in the framework region. Alternative positions will be explored in new MD simulations, searching for variants with improved affinities, to be tested experimentally.
Prediction of B/T cell epitopes by improved in silico methods and doing the immunogenicity testing in vitro. Currently no generally applicable procedure is available for assessing the expected immunogenicity of therapeutic antibodies combined with prediction of molecular function, i.e. binding affinity. Human germline frameworks can be considered as non-immunogenic mainly because of different immunotolerance mechanisms. However, variants with a significant amount of murine residues have improved binding properties (Mader et al. 2010). In these regions the sequence can deviate from germline encoded antibody sequences generating novel epitopes that may trigger possible immunogenic reactions. We propose that currently available in silico prediction tools to predict immunogenicity may be used to identify relevant positions for backmutations in (partially humanized) variants. These may be prioritized based on molecular simulations and subsequently expressed and tested to establish humanized Ab2/3H6 variants with minimal expected immunogenicity but fully functional binding capacities.
Mader A. and R. Kunert (2012) Evaluation of the potency of the anti-idiotypic antibody Ab2/3H6 mimicking gp41 as an HIV-1 vaccine in a rabbit prime/boost study. PLoS One; 7(6): e39063.
Kunert, R. and A. Mader (2011) Anti-idiotypic antibody Ab2/3H6 mimicking gp41: a potential HIV-1 vaccine? BMC Proceedings, 5; ISSN 1753-6561
de Ruiter, A., A. Mader, R. Kunert and C. Oostenbrink (2011) Molecular Simulations to Rationalize Humanized Ab2/3H6 Activity. Aust J Chem.; 64(7): 900-909.
Mader, A and R. Kunert (2010) Humanization strategies for an anti-idiotypic antibody mimicking HIV-1 gp41. Protein Eng Des Sel; 23(12):947-954
Gach, J.S., H. Quendler, S. Strobach, H. Katinger and R. Kunert (2008) Structural analysis and in vivo administration of an anti-idiotypic antibody against mAb 2F5. Mol Immunol; 45: 1027–1034.
Bryson, S., J.P. Julien, D.E. Isenman, R. Kunert, H. Katinger and E.F. Pai (2008) Crystal Structure of the Complex between the Fab fragment of the Cross-Neutralizing Anti-HIV-1 Antibody 2F5 and the Fab Fragment of Its Anti-idiotypic Antibody 3H6. J. Mol. Biol.; 382(4): 910–919.
