Isolation and functional characterization of Nicotiana benthamiana serine proteinases involved in the degradation of recombinant proteins
SUPERVISOR: Lukas MACH
Background.
Several studies have demonstrated that plants are suitable hosts for the production of protein therapeutics (de Muynck et al., 2010). Unfortunately, the production of recombinant proteins in plants is frequently hampered by unwanted proteolysis (Benchabane et al., 2008). For recombinant monoclonal antibodies, the generation of substantial amounts of Fab- and Fc-like degradation products has been observed in different plant species (Doran, 2006). This is also the case in Nicotiana benthamiana, a tobacco-related plant species widely used for recombinant protein production (Villani et al., 2009). We have now obtained evidence that in planta fragmentation of antibodies involves the action of serine proteinases (Niemer et al., 2014). However, the N. benthamiana representatives of this family of proteolytic enzymes are so far only poorly characterized (Goulet et al., 2012).
Aims and methods.
This project aims at the identification and detailed characterization of N. benthamiana serine proteinases capable of degrading monoclonal antibodies. Special attention will be given to those enzymes, which reside within the apoplast since antibody degradation of recombinant proteins occurs largely in this compartment (Niemer et al., 2014). Recent progress in the development of activity-based probes for serine proteinases has allowed the detection of active species of these enzymes in plant extracts (van der Hoorn and Kaiser, 2012). A biotin-tagged version of such a functional probe will be used for the selective labelling of serine proteinases present in N. benthamiana apoplastic fluid. After isolation with immobilized streptavidin, the captured proteins will then be identified by mass spectrometry as previously reported by us for another class of proteinases (Gu et al., 2012). Exploiting the availability of the recently elucidated N. benthamiana genome sequence, the respective cDNAs will be cloned by RT-PCR using primers based on the peptide sequences obtained by mass spectrometry (ALTMANN) and then expressed in insect cells (Liebminger et al., 2013). The recombinant proteinases will be purified exploiting exogenously added affinity tags and then analysed for their capacity to degrade the particularly vulnerable monoclonal anti-HIV-1 antibodies 2F5 and PG9. The cleavage products obtained upon digestion of 2F5 and PG9 with the recombinant proteinases will then be characterized by N-terminal sequencing and compared to those generated by incubation of the antibodies with unfractionated apoplastic fluid (Niemer et al., 2014).
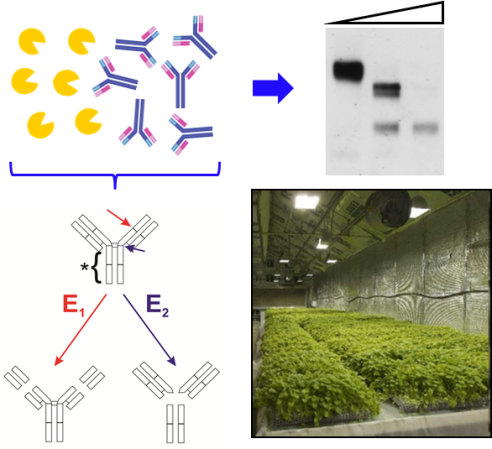
The second major aim of this project is the generation of N. benthamiana lines with reduced activities of antibody-degrading serine proteinases. We will use an RNA interference (RNAi) strategy based on agrobacterium-mediated transfection for silencing of these genes, initially in a transient manner. Stable RNAi lines will then be generated by means of the approach previously used for the successful knock-down of different glycosylation enzymes (Strasser et al., 2008). To assess the effects of down-regulation of apoplastic serine proteinase activities on recombinant protein integrity and yield, the proteolysis-sensitive antibodies 2F5 and PG9 will then be transiently co-expressed in the respective plants (STEINKELLNER). The antibodies will be purified by affinity chromatography and characterized by electrophoresis, mass spectrometry and N-terminal sequencing. Their structural integrity will be evaluated by size-exclusion chromatography and circular-dichroism spectroscopy (OBINGER). Assessment of the antigen-binding properties of the plant-derived antibodies will be performed by ELISA and biolayer interferometry in comparison to antibodies produced in mammalian cells (KUNERT).
Benchabane, M., Goulet, C., Rivard, D., Faye, L., Gomord, V., Michaud, D. (2008) Preventing unintended proteolysis in plant protein biofactories. Plant Biotechnol. J. 6, 633-648
de Muynck., B., Navarre, C., Boutry, M. (2010) Production of antibodies in plants: status after twenty years. Plant Biotechnol. J. 8, 529-563
Doran, P. M. (2006) Foreign protein degradation and instability in plants and plant tissue cultures. Trends Biotechnol. 24, 426-432
Goulet, C., Khalf, M., Sainsbury, F., D'Aoust, M.-A., Michaud, D. (2012) A protease activity-depleted environment for heterologous proteins migrating towards the leaf cell apoplast. Plant Biotechnol. J. 10, 83-94
Gu, C., Shabab, M., Strasser, R., Wolters, P. J., Shindo, T., Niemer, M., Kaschani, F., Mach, L., van der Hoorn, R. A. (2012) Post-translational regulation and trafficking of the granulin-containing protease RD21 of Arabidopsis thaliana. PLoS One 7:e32422
Liebminger, E., Grass, J., Altmann, F., Mach, L., Strasser, R. (2013) Characterizing the link between glycosylation state and enzymatic activity of the endo-β1,4-glucanase KORRIGAN1 from Arabidopsis thaliana. J. Biol. Chem. 288, 22270-22280
Niemer, M., Mehofer, U., Torres Acosta, J. A., Verdianz, M., Henkel, T., Loos, A., Strasser, R., Maresch, D., Rademacher, T., Steinkellner, H., Mach, L. (2014) The human anti-HIV antibodies 2F5, 2G12 and PG9 differ in their susceptibility to proteolytic degradation: Downregulation of endogenous serine and cysteine proteinase activities could improve antibody production in plant-based expression platforms. Biotechnol. J. 9, 493-500
Strasser, R., Stadlmann, J., Schaehs, M., Stiegler, G., Quendler, H., Mach, L., Gloessl, J., Weterings, K., Pabst, M., Steinkellner, H. (2008) Generation of glyco-engineered Nicotiana benthamiana for the production of monoclonal antibodies with a homogeneous human-like N-glycan structure. Plant Biotechnol. J. 6, 392-402
van der Hoorn, R. A., Kaiser, M. (2012) Probes for activity-based profiling of plant proteases. Physiol. Plant. 145, 18-27
Villani, M. E., Morgun, B., Brunetti, P., Marusic, C., Lombardi, R., Pisoni, I., Bacci, C., Desiderio, A., Benvenuto, E., Donini, M. (2009) Plant pharming of a full-sized, tumour-targeting antibody using different expression strategies. Plant Biotechnol. J. 7, 59-72
