Divergent evolution of the CDE superfamily
PRINCIPAL INVESTIGATOR: CHRISTIAN OBINGER
Background.
Heme b containing chlorite dismutases (Clds) catalyze the decomposition of chlorite (ClO2-) into chloride (Cl-) and molecular oxygen (O2). During turnover a covalent oxygen-oxygen bond is formed, a unique biochemical reaction so far only found in the water-splitting manganese complex of photosystem II of oxygenic organisms. Genes encoding Clds are present in numerous bacterial and archaeal phyla thus representing rather ancient sequences. Recently, our group together with DJINOVIC-CARUGO succeeded in obtaining high-resolution X-ray crystal structures together with spectral, redox and mechanistic data of functional Clds (Cld from Nitrospira defluvii (NdCld) and Cld from the cyanobacterium Cyanothece (CCld)). NdCld and CCld are representatives of the two main (functional) Cld clades that differ in subunit architecture and oligomeric structure but exhibit very similar chlorite degradation kinetics (Hofbauer et al. 2012, 2014a, 2014b). All functional (i.e. chlorite degrading) Clds have a fully conserved distal arginine, which participates in chlorite degradation.
Phylogenetic analysis and structural homologies clearly demonstrate connections of functional Clds to two other large, ancient, and functionally different protein families. One includes Cld-like proteins (that lack the distal arginine and cannot degrade chlorite), also called HemQ, found in bacteria and archaea. The physiological role of Cld-like proteins is unknown and hypotheses about their role(s) include participation in biosynthesis of heme or antibiotics. The second protein family is comprised of recently discovered dye-decolorizing heme b-containing peroxidases (DyPs, with bacterial representatives also called EfeBs). They exhibit a novel active site architecture, which is different compared to all so far described heme peroxidase families and contains a distal arginine/aspartate pair. DyPs oxidize bulky molecules (e.g. anthraquinone derivatives) as well as artificial electron donors (e.g. ABTS, azo-dyes). They are found in prokaryotes and – in contrast to Clds and Cld-like proteins – also in eukaryotes, mainly fungi.
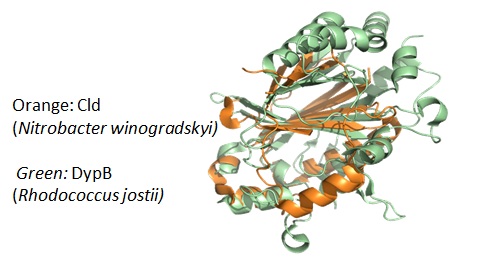
Clds, Cld-like proteins (HemQ), DyPs and EfeBs comprise the CDE-superfamily that has evolved from a common ancestor (Goblirsch et al., 2011; Hofbauer et al., 2014c). The present project aims at understanding the functional diversification of this superfamily during evolution by analysis of the structural and functional relationship of current Clds and DyPs as well as reconstructed ancestral Clds.
Aims and methods.
Despite sequence and structure homology Clds have negligible peroxidase activity, whereas DyPs are unable to degrade chlorite. Based on the established expression in E. coli and the availability of high-resolution structures, NdCld and CCld will be used as model enzymes that will be engineered for interconversion to DyPs. Additionally, we aim at implementation of chlorite degradation activity into DyP-A from Rhodococcus jostii. Designed mutants will be analysed with respect to spectral, redox and kinetic properties and compared with the wild-type enzymes. Selected variants will be characterized in more detail including resonance Raman-(SMULEVICH) and continuous-wave and pulsed electronic paramagnetic resonance (VAN DOORSLAER) spectroscopy, multi-mixing stopped-flow spectroscopy, spectroelectrochemistry (BATTISTUZZI), mass spectrometry (ALTMANN and WILSON) and X-ray crystallography (DJINOVIC-CARUGO). Those methods will help to elucidate the role (i) of distal residues in catalysis, (ii) of inner- and outer-sphere ligands and H-bonding networks in interconversion of enzyme intermediates and redox thermodynamics and (iii) substrate binding and oxidation site(s) in the respective mutants and, finally, in wild-type Clds and bacterial DyPs.
Furthermore, it is planned to perform in silico reconstruction of ancestral Clds by phylogenetic inference. Ancestral protein reconstruction offers additional options to study interrelationships of function and structure of proteins and can reveal synergistic effects of multiple mutations (Harms et al., 2010). Based on critical inspection of obtained sequences in relation to structural and functional data of wild-type and engineered Clds and DyPs, selected synthetic genes will be expressed in E. coli and the respective proteins biochemically characterized. By both protein engineering and descending the phylogeny stepwise toward the common ancestor of Clds, we aim at a better understanding of the divergent evolution of the CDE superfamily.
Goblirsch, B.R., Kurker, R.C., Streit, B.R., Wilmot, C.M., Dubois, J.L. (2011) Chlorite dismutases, DyPs, and EfeB: 3 microbial heme enzyme families comprise the CDE structural superfamily. J. Mol. Biol. 408, 379-398
Harms, M.J., Thornton, J.W. (2010) Analyzing protein structure and function and using ancestral gene reconstruction. Curr. Opin. Struct. Biol. 20, 360-366
Hofbauer, S., Bellei, M., Suendermann, A., Pirker, K.F., Hagmueler, A., Mlynek, G., Kostan, J., Daims, H., Furtmueller, P.G., Djinović-Carugo, K., Oostenbrink, C., Battistuzzi, G., Obinger, C. (2012) Redox thermodynamics of high-spin and low-spin forms of chlorite dismutases with diverse subunit and oligomeric structures. Biochemistry 51, 9501-9512
Hofbauer, S., Gruber, C., Pirker, K.F., Suendermann, A., Schaffner, I., Jakopitsch, C., Oostenbrink, C., Furtmueller, P.G., Obinger, C. (2014a) Transiently produced hypochlorite is responsible for the irreversible inhibition of chlorite dismutase. Biochemistry 53, 3145–3157
Hofbauer, S., Gysel, K., Bellei, M., Pirker, K.F., Hagmueller, A., Schaffner, I., Mlynek, G., Kostan, J., Daims, H., Furtmueller, P.G., Battistuzzi, G., Djinovic-Carugo, K., Obinger, C. (2014b) Manipulating conserved heme cavity residues of chlorite dismutase: effect on structure, redox chemistry and reactivity. Biochemistry 53, 77-89
Hofbauer, S., Schaffner, I., Furtmueller, P.G., Obinger, C. (2014c) Chlorite degradation, oxygen generation and broad peroxidase specificity – A novel heme enzyme family for use in bioremediation and biosynthesis. Biotechnol. J. 9, 461-473
