Modelling and simulations of glycosylated proteins
SUPERVISOR: CHRIS OOSTENBRINK
Background.
Computer simulation of biomolecular complexes has developed into a powerful tool, complementary to experimental exploration (Karplus and McCammon, 2002; Van Gunsteren et al., 2006). By zooming in on the atomic level, complex biomolecular systems can be described at a time and space resolution often unattainable by experiments, thereby leading to significant insights into the workings of the molecular machinery of the cell. For proteins and nucleic acids, computer simulations are readily applied. In spite of their relevance, post-translational modifications are strongly underrepresented in such simulations. We have recently provided interaction parameters for a large set of post-translational modifications (Petrov et al., 2013). In particular, glycosylation is only rarely taken into account in computer models, due to the heterogeneity of the glycosylation and lack of appropriate three dimensional models. Significant improvements on the description of atomic interactions in carbohydrates have been described recently (Lins andHünenberger, 2005; Hansen and Hünenberger, 2011), opening the way to a more extensive molecular mechanics based modelling of these molecular structures. Recently, we have extensively studied the interactions of proteins with small carbohydrates (Garate and Oostenbrink, 2013; Graf et al., 2013).
The determination of the exact molecular structure of glycosylated proteins is often a challenge by itself. Even if the molecular structure has been resolved, complete 3D molecular structures are only rarely resolved in NMR or X-ray crystallography. Glycan structures may be modeled onto available deglycosylated protein structures, but the intrinsic flexibility of the links between individual carbohydrates leads to a multitude of possible structures. Molecular simulations may converge many of these to the most prominent conformation while intrinsically, the relevant ensemble of glycan conformations will need to be sampled.
Aims and methods.
Tools will be developed to generate relevant glycan structures in the absence of experimental structural data. Biomolecular polymeric structures like proteins, DNA and RNA are characterized by well-defined three-dimensional structures, which can be classified in terms of secondary or tertiary structure. We have recently established a new classification scheme for the analysis of protein and polynucleotide structures (Nagy and Oostenbrink, 2014a,b). For carbohydrates, much less experimental structural data is available and to date no universal definition of secondary structure for carbohydrates has been proposed. We will perform a thorough analysis of available 3D glycan structure, based on which a definition of the most prominent dihedral angle structures and possibly of recurring structural motifs may be established.
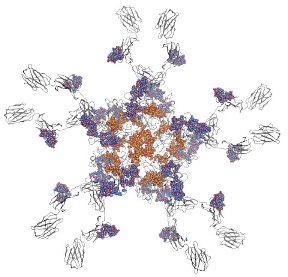
This information will be used to construct possible models of additional glycan structures, which will subsequently be optimized through the development of enhanced sampling tools that search the relevant conformational space of complex molecular structures (Peric-Hassler et al., 2010). Once models of glycan structures may reliably be generated, the effect of these on the structure, dynamics and function of the related proteins will be studied through computer simulations. To this effect, different protein constructs will be simulated, either in the absence of glycan structures, or with different glycosylation patterns. A close relationship with experimental work on controlled glycosylation in plant systems should be established such that the simulations may aid in the rationalization of experimental observations on differently glycosylated proteins. Specific glycosylation has been observed for different proteins, or even within a single protein structure (Loos et al., 2014). An explanation for this fact may reside in steric factors determining what kind of glycan structure may be placed at specific locations, or by the specific accessibility of glycosyltransferases (Thaysen-Andersen and Packer, 2012). The distinct glycosylation patterns at specific sites of IgMs as well as the structure and dynamics of selected glycan-modifying enzymes and novel glycans of interest within BioToP will be studied, thereby enabling intelligent protein engineering or rationalization of binding properties.
Garate, J.A., Oostenbrink, C. (2013) Lipid A from lipopolysaccharide recognition: Structure, dynamics and cooperativity by molecular dynamics simulations. Proteins 81, 658-674
Graf, M.M.H., Bren, U., Haltrich, D., Oostenbrink, C. (2013) Molecular dynamics simulations give insight into D-glucose dioxidation at C2 and C3 by Agaricus meleagris pyranose dehydrogenase. J. Comp. Aid. Mol. Des. 27, 295-304
Hansen, H.S., Huenenberger, P.H. (2011) A reoptimized GROMOS force field for hexopyranose-based carbohydrates accounting for the relative free energies of ring conformers, anomers, epimers, hydroxymethyl rotamers and glycosidic linkage conformers. J. Comput. Chem. 32, 998–1032
Karplus, M., McCammon, J.A. (2002) Molecular dynamics simulations of biomolecules. Nature Struct. Biol. 9, 646-652
Lins, R.D., Huenenberger, P.H. (2005) A new GROMOS parameter set for hexopyranose-based cardohydrates. J. Comput. Chem. 26, 1400-1412
Loos, A., Gruber, C., Altmann, F., Mehofer, U., Hensel, F., Grandits, M., Oostenbrink, C., Stadlmayr, G., Furtmueller, P.G., Steinkellner, H. (2014) Expression and glycoengineering of functionally active heteromultimeric IgM in plants. Proc. Natl. Acad. Sci. USA 111, 6263-6268
Nagy, G., Oostenbrink, C. (2014a) Dihedral-based segment identification and classification of biopolymers I: Proteins. J. Chem. Inf. Model. 54, 266–277
Nagy, G., Oostenbrink, C. (2014b) Dihedral-based segment identification and classification of biopolymers II: Polynucleotides. J. Chem. Inf. Model. 54, 278–288
Peric-Hassler, L., Hansen, H.S., Baron, R., and Hünenberger, P H. (2010) Conformational properties of glycose-based disaccharides investigated using molecular dynamics simulations with local elevation umbrella sampling. Carbohydr. Res. 345, 1781–1801
Petrov, D., Margreitter, C., Grandits, M., Oostenbrink, C., Zagrovic, B. (2013) A systematic framework for molecular dynamics simulations of protein post-translational modifications. PLOS Comput. Biol. 9:e1003154
Thaysen-Andersen, M., Packer, N.H. (2012) Site-specific glycoproteomics confirms that protein structure dictates formation of N-glycan type, core fucosylation and branching. Glycobiology 22, 1440-1452
Van Gunsteren, W.F., Bakowies, D., Baron, R., Chandrasekhar, I., Christen, M., Daura, X., Gee, P., Geerke, D., Glaettli, A., Huenenberger, P.H., Kastenholz, M.A., Oostenbrink, C., Schenk, M., Trzesniak, D., van der Vegt, N.F.A., and Yu, H.B. (2006) Biomolecular modeling: goals, problems, perspectives. Angew. Chem. Int. Ed. 45, 4064–4092
