Synthetic biology tools to control expression of multiple genes in a pathway
SUPERVISOR: NICOLE BORTH
Background.
According to theoretical calculations, current production processes of recombinant proteins take advantage of at best 25% of the cellular capacity available for such production (Hefzi et al., 2016). The reasons for this lack of targeted resource usage are not clear.
According to model predictions, the most successful strategies would be genetic engineering approaches, however, to take systematic advantage of these, rational design strategies for multiplexed engineering approaches are desperately needed. These would have to target the regulation of entire pathways, rather than aiming for the traditional overexpression of individual genes. Such modifications of the expression levels of a connected set of endogenous genes could shift the flux of native pathways towards improved phenotypes, but would also require extensive recombination and changes in the endogenous network. To identify the best combinations and synergies of expression levels, it will be necessary to fine-tune the expression rate of multiple endogenous genes simultaneously, but individually and independently. Traditional genetic engineering approaches rely on stable overexpression or knock-out/knock-down of target genes (Hansen et al., 2017), thus considering only on/off states of individual genes, ignoring the full dynamics and regulation of pathways which may change in dependence of the culture state and process conditions.
Such tools are currently emerging in Synthetic Biology research (Chen and Ho, 2017), but may have to be adapted and further developed to represent the entire dynamic range required and to achieve compatibility with the CHO system. Ideally, such control tools would enable the post-engineering adjustment of gene expression levels depending on the specific task required at any time by the cell line (such as different protein categories that may require different processing steps) or the cultivation process (during subcloning, expansion or bioprocess production). The focus in this project will therefore lie on the development of control elements that enable externally controllable, inducible or responsive (for instance to culture conditions or cellular state) gene expression.
Aims and methods.
The project aims to establish a set of responsive control elements that define gene expression strength according to external signals (such as added chemicals or inducers, or physical signals such as temperature) or in response to intracellular signals that mark the state of the cell (such as growth rate, nutrient supply or similar).
At least four independently controllable elements are required to enable pathway engineering (such that each gene can be independently and in parallel regulated and the optimal settings be adjusted for given process conditions). The following responsive control systems will be investigated:
Inducible CRISPR/Cas9 systems using activation or inactivation domains in combination with degrons or chemically induced heterodimerization (Natsume and Kanemaki, 2017)
siRNAs/miRNAs, where the biological mechanism of regulation is well studied. By modulating RNA interference through small molecule interaction and varying the number of target sites on the mRNA, gene expression can be controlled artificially (Jadhav et al., 2013)
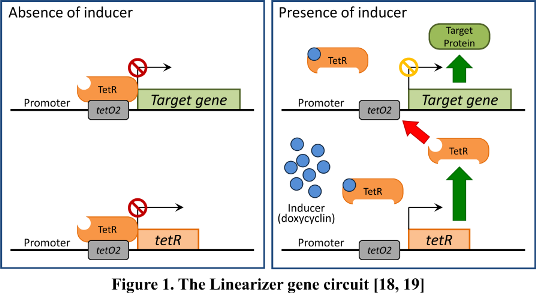
Linearizer circuits which respond to externally added molecules and which we will use to replace endogenous promoters of identified target genes by CRISPR/Cas9 (Nevozhay et al., 2013)
Riboswitches are flexible RNA structures that can control protein production through RNA cleavage (Chen et al., 2010) and splicing (Li and Breaker, 2013), thus offering an even wider range of control mechanisms beyond transcription.
Once a set of such control elements is identified and tested in CHO, we will select a suitable pathway, such as glycosylation, and set important genes within this pathway under control. By fine-tuning the respective expression level of these genes one would expect to see different glycan structures on the expressed model protein.
Collaborations within this thesis will include ALTMANN (glycoanalysis) and STRASSER (protein glycosylation).
Hefzi, H., Ang, K.S., Hanscho, M., Bordbar, A., …, Borth, N., Lee, D.Y., Lewis, N.E. (2016) A consensus genome scale reconstruction of Chinese Hamster Ovary cell metabolism. Cell Sys. 23, 3, 434-443. doi: 10.1016/j.cels.2016.10.020
Hansen, H.G., Pristovsek, N., Kildegaard, H.F., Lee, G.M. (2017) Improving the secretory capacity of Chinese hamster ovary cells by ectopic expression of effector genes: lessons learned and future directions. Biotechn. Adv. 35, 64-76. doi: 10.1016/ j.biotechadv.2016.11.008
Chen, Y.Y. and Ho, P. (2017) Mammalian synthetic biology in the age of genome editing and personalized medicine. Curr. Opin. Chem. Biol. 40, 57-64. doi: 10.1016/ j.cbpa.2017.06.003
Natsume, T. and Kanemaki, M.T. (2017) Conditional degrons for controlling protein expression at the protein level. Ann. Rev. Genetics 51, 83-102. doi: 10.1146/annurev-genet-120116-024656
Jadhav, V., Hackl, M., Druz, A., Shridhar, S., Chung, C-Y., Heffner, K.M., Kreil, D.P., Betenbaugh, M., Shiloach, J., Barron, N., Grillari, J., Borth, N. (2013) CHO microRNA engineering is growing up: Recent successes and future challenges. Biotechn. Adv. 31, 1501-1513. doi: 10.1016/j.biotechadv.2013.07.007
Nevozhay, D., Zal, T., Balazsi, G. (2013) Transferring a synthetic gene circuit from yeast to mammalian cells. Nat. Commun. 4, 1451. doi: 10.1038/ncomms2471
Chen, Y.Y., Jensen, M.C., Smolke, C.D. (2010) Genetic control of mammalian T-cell proliferation with synthetic RNA regulatory systems. Proc. Natl. Acad. Sci. USA 107, 8531-8536. doi: 10.1073/pnas.1001721107
Li, S. and Breaker, R.R. (2013) Eukaryotic TPP riboswitch regulation of alternative splicing involving long-distance base pairing. Nucl. Acids Res. 41, 3022–3031. doi: 10.1093/nar/gkt057
