Mechanisms of control of endogenous gene expression
SUPERVISOR: NICOLE BORTH
Background.
Recent work in our group has highlighted the importance of long non-coding RNAs in regulation of gene transcription in CHO cells. During a batch culture, we found 14,547 coding genes expressed in CHO cells, of which 1,397 genes were differentially expressed. At the same time, 42,177 long non-coding genes (lncRNAs) were expressed, 2,899 of these differentially. Such abundantly transcribed genes must have an important cellular function.
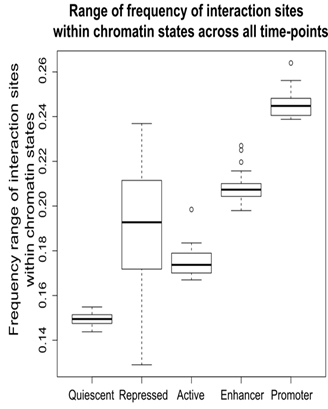
One proposed mechanisms of action of lncRNAs is the formation of DNA-DNA-RNA triplexes which can lead to both upregulation and attenuation of coding gene expression. Indeed, we found that the majority of predicted triplex-forming interactions of expressed lncRNAs were within promoter regions, followed by enhancers and regions of the genome that are actively transcribed.
For the differentially expressed lncRNAs, all predicted interactive coding genes were also differentially expressed, both in a corresponding or in a counterregulated way. Interestingly, the genes with the highest fold change over the batch culture had the lowest presence of histone marks that characterize active transcription and the highest number of triplex forming interactions. It thus seems that lncRNAs play an important role in rapid control of gene expression level and thus are also promising tools that might be used for manipulation of gene expression in the native genomic context.
Aims and methods.
To obtain a better understanding of the mechanisms underlying regulation of gene expression levels by lncRNAs, such as attraction or blocking of transcription factors to the site of triplex formation, we will study the sequences of lncRNAs expressed in CHO cells and their effect on target gene expression, using a combination of bioinformatic and wet-lab approaches. In essence, lncRNA sequences that have been identified to either enhance or repress gene expression will be tested on artificial constructs in-vitro, to verify their effect.
In addition, network analysis of the different genes and their targets, as based on co-expression and interaction between regulated genes will be analysed to obtain a deeper understanding of phenotypic control.
Collaborations within this thesis will include ZANGHELLINI (metabolic modelling).
Hefzi, H., Ang, K.S., Hanscho, M., Bordbar, A., …, Borth, N., Lee, D.Y., Lewis, N.E. (2016) A consensus genome scale reconstruction of Chinese Hamster Ovary cell metabolism. Cell Sys. 23, 3, 434-443. doi: 10.1016/j.cels.2016.10.020
Zucchelli, S., Fasolo, F., Russo, R., Cimatti, L., Patrucco, L., Takahashi, H., Jones, M.H., Santoro, C., Sblattero, D., Cotella, D., Persichetti, F., Carninci, P., Gustincich, S. (2015) SINEUPs are modular antisense long non-coding RNAs that increase synthesis of target proteins in cells. Front. Cell. Neurosci. 9, 174. doi: 10.3389/fncel.2015.00174
Zucchelli, S., Patrucco, L., Persichetti, F., Gustincich, S., Cotella, D. (2016) Engineering translation in mammalian cell factories to increase protein yield: the unexpected use of long non-coding SINEUP RNAs. Comp. Struct. Biotechn. J. 14, 404-410. doi:10.1016/j.csbj.2016.10.004
