Protein solubility in buffers with kosmotropic salts and polyols
SUPERVISOR: ALOIS JUNGBAUER
Background.
The solubility of proteins is enhanced by low concentrations of kosmotropic salts (as described by the Hofmeister series), amino acids, and polyols. These compounds also increase the stability of the proteins. Often this is explained by preferential interaction theory, which do not a priori predict if these additives increase stability and if the compound is accumulated in the vicinity of the protein surface. Moreover a single protein concentration at dilute conditions is represented by this theory.
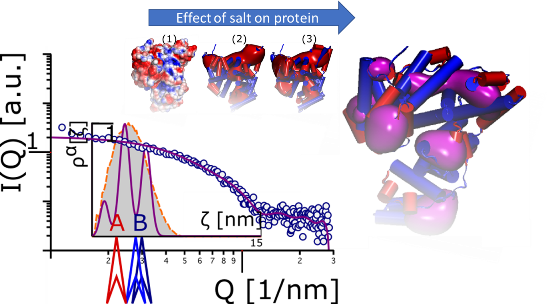
For a holistic understanding of protein solubility it is necessary to understand effects along the entire phase diagram of a protein. Furthermore, for formulation (Mueller et al., 2013a, b), precipitation, and crystallization (Huettmann et al., 2015), of proteins and other biologics the effect of additives on solubility is very important. Phase diagrams are measured by high throughput methods and the solubility line is approximated by a simple log-linear equation yield in a salting constant or by the extended Debye Hueckel theory, which also does not cover the entire concentration range.
Aims and methods.
The aim of this thesis is to find out if additives (sodium chloride, ammonium sulphate, sucrose and trehalose) increasing the protein solubility are accumulating close to the surface of proteins and if this is in line with the preferential interaction theory.
The research hypothesis is as follows: When the additive increases solubility the additive accumulates close to the surface of protein. This is in part in contradiction to the preferential interaction theory. We will solve this problem with the following methodology: as model protein we use small proteins such as lysozyme, cytochrome c, basic fibroblast growth factor and tumour necrosis factor. The effect of the additive on solubilisation will be tested by high throughput screening in microtiter plates to get the solubility curve for increasing additive concentration. The solubility curve will be approximated numerically. The additive concentration close to the surface concentration will be determined by isotope labelled additives by NMR. The size of the protein with increasing additive concentration will be determined by dynamic light scattering, multi-angel light scattering and SAXS. The radius of the protein with and without the additive will be calculate and compared to simulation.
Collaborations within this thesis will include OOSTENBRINK (simulation of protein structure and size).
Huettmann, H., Zich, S., Berkemeyer, M., Buchinger, W., Jungbauer, A. (2015) Design of industrial crystallization of interferon gamma: Phase diagrams and solubility curves. Chem. Eng. Sci. 126, 341-348. doi: 10.1016/j.ces.2014.12.018
Mueller, M., Loh, M.Q.T., Tscheliessnig, R., Tee, D.H.Y., Tan, E., Bardor, M., Jungbauer, A. (2013a) Liquid formulations for stabilizing IgMs during physical stress and long-term storage. Pharm. Res. 30,735-750. doi: 10.1007/s11095-012-0914-2
Mueller, M., Loh, M.Q.T., Tee, D.H.Y., Yang, Y., Jungbauer, A. (2013b) Liquid formulations for long-term storage of monoclonal IgGs. Appl. Biochem. Biotechnol. 169, 1431-1448. doi: 10.1007/s12010-012-0084-z
