Substrate specificity in carbohydrate oxidoreductases
PRINCIPAL INVESTIGATOR: ROLAND LUDWIG
Background.
Blood glucose meters and wearable continuous glucose measurement systems (CGMS) are important tools for people with diabetes for the maximization of their welfare. Precisely dosed insulin injections are a prerequisite for wellbeing, and individually optimized treatment strategies and adjustments should be based on reliable blood glucose measurements. Glucose meters and CGMS depend on the substrate specificity of their enzymes for accurate, precise and reliable measurements. The most frequently employed biorecognition elements are FAD-dependent glucose oxidase and glucose dehydrogenase from the glucose-methanol-choline (GMC)-oxidoreductase family. Despite their long history of use, the catalytic and electron transfer properties of these enzymes still compromise biosensor design by their dependence on oxygen or redox mediators, their susceptibility to interfering natural and pharmaceutical substances, and their insufficient substrate specificity (Ramos et al., 2008; Frias et al., 2010). These insufficiencies are the driving factors for the development of third-generation glucose biosensors, which employ enzymes capable of direct electron transfer (DET) at a low working electrode potential to reduce interferences. Cellobiose dehydrogenase (CDH) is such a DET-capable oxidoreductase and a promising biorecognition element for third-generation biosensors (Tasca et al., 2011; Felice et al., 2013).
CDH is a typical GMC oxidoreductase, but comprises an additional cytochrome domain that serves as a build-in redox mediator (Tan et al., 2015). The general principles of substrate recognition and catalysis by GMC oxidoreductase have been investigated by collaborators employing X-ray diffraction (Hallberg et al., 2002; Kujawa et al., 2006; Tan et al., 2013) and fast kinetic methods (Wongnate and Chaiyen, 2013). However, experimental studies in our group to change the substrate specificity of carbohydrate converting GMC oxidoreductases for biocatalytic applications were aimed at increased substrate turnover rates (Spadiut et al., 2009) and resulted in biocatalysts showing substrate specificity not adequate for biosensors. In this project, a systematic approach will be applied to elucidate the mechanistic determinants of glucose binding both in glucose oxidase and glucose dehydrogenase, which will serve as a base to redesign the cellobiose-binding site of CDH to a glucose-binding site. In contrast to earlier attempts, which were based on single-point mutations, multiple parallel amino acid exchanges will be performed to efficiently rebuild CDH's binding site for highly specific glucose binding. Kinetic characterisation of CDH variants will be used to experimentally quantify the effects of mutations and verify or falsify the hypotheses behind the amino acid exchanges.
Aims and methods.
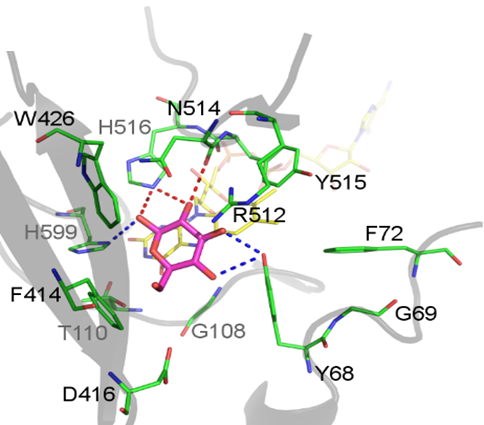
In this thesis project, we aim to elucidate the general principles of substrate recognition and catalysis in carbohydrate-converting GMC oxidoreductases and apply the principles to generate a CDH with high glucose specificity as a biorecognition element for a DET-based glucose biosensor. The active sites of glucose oxidases, glucose dehydrogenases and CDHs will be computationally investigated to determine the mechanistic factors for mono- and disaccharide binding. Docking and molecular dynamics simulations will be used to evaluate the binding contribution of individual amino acid residues and the active-site structure (OOSTENBRINK). Using the same methodology, redesigned CDH active sites will be evaluated in silico, before constructing and producing enzyme variants.
CDHs with redesigned active sites will be constructed by site-directed mutagenesis introducing one or multiple mutations at a time. If several positions have to be targeted, multi-site-directed mutagenesis will be applied. This technique will also be used to generate multiple randomized mutations on selected hotspots. Libraries of enzyme variants will be screened by photometric high-throughput assays. Expression of mutated cdh genes will be performed in the yeast P. pastoris. Production of CDH variants will be performed in shaking flasks, or if necessary in bioreactors. A three-step purification procedure for obtaining homogeneous CDH is established (Harreither et al., 2011).
The specific CDH variants that may have an increased substrate specificity for glucose and a decreased specificity for disaccharides will be tested by steady-state spectrophotometry to determine KM, kcat, and the specificity constant. Using stopped-flow photometry, the individual half-reactions will be investigated (HALTRICH and OBINGER). The reductive-half reaction will be studied for the analyte glucose, the interfering saccharides maltose and lactose, and the natural substrate cellobiose. Binding and dissociation rate constants will be determined to evaluate the effect of changes in the binding site. Cyclic voltammetry and chronoamperometry will be used to evaluate the heterogeneous kinetics of electrode-bound CDH variants and compare their performance with currently used glucose biosensors.
Collaborations within this thesis include OOSTENBRINK (molecular dynamics), HALTRICH (fast kinetics) and OBINGER (enzymatic mechanisms).
Felice, A.K.G., Sygmund, C., Harreither, W., Kittl, R., Gorton, L., Ludwig, R. (2013) Substrate specificity and interferences of a direct-electron-transfer-based glucose biosensor. J. Diabet. Sci. Technol. 7, 669-677. doi: 10.1177/193229681300700312
Frias, J. P., Lim, C. G., Ellison, J. M., Montandon, C. M. (2010) Review of adverse events associated with false glucose readings measured by GDH-PQQ–based glucose test strips in the presence of interfering sugars. Diabetes Care 33, 728-729. doi: 10.2337/dc09-1822
Hallberg, M.B., Henriksson, G., Pettersson, G., Divne, C. (2002) Crystal structure of the flavoprotein domain of the extracellular flavocytochrome cellobiose dehydrogenase. J. Mol. Biol. 315, 421-434. doi: 10.1006/jmbi.2001.5246
Harreither, W., Sygmund, C., Augustin, M., Narciso, M., Rabinovich, M.L., Gorton, L., Haltrich, D., Ludwig, R. (2001) Catalytic properties and classification of cellobiose dehydrogenases from ascomycetes. Appl. Environ. Microbiol. 77, 1804-1815. doi: 10.1128/AEM.02052-10
Kujawa, M., Ebner, H., Leitner, C., Hallberg, B.M., Prongjit, M., Sucharitakul, J., Ludwig, R., Rudsander, U., Peterbauer, C., Chaiyen, P., Haltrich, D., Divne, C. (2006) Structural basis for substrate binding and regioselective oxidation of monosaccharides at C3 by pyranose 2-oxidase. J. Biol. Chem. 281, 35104-35115. doi: 10.1074/jbc.M604718200
Ramos, R., González, T., Fulladosa, X., Castelao, A.M., Grinyó, J.M. (2008) False hyperglycemies in diabetic patients using icodextrin in peritoneal dialysis. Dial. Transplant. 37, 323-327. doi: 10.1002/dat.20229
Spadiut, O., Radakovits, K., Pisanelli, I., Salaheddin, C., Yamabhai, M., Tan, T.-C., Divne, C., Haltrich, D. (2009) A thermostable triple mutant of pyranose 2-oxidase from Trametes multicolor with improved properties for biotechnological applications. Biotechnol. J. 4, 525-534. doi: 10.1002/biot.200800260
Tan, T.C., Spadiut, O., Wongnate, T., Sucharitakul, J., Krondorfer, I., Sygmund, C., Haltrich, D., Chaiyen, P., Peterbauer, C.K., Divne, C. (2013) The 1.6 Å crystal structure of pyranose dehydrogenase from Agaricus meleagris rationalizes substrate specificity and reveals a flavin intermediate. PLoS One 8, e53567. doi: 10.1371/journal.pone.0053567
Tan, T.-C., Kracher, D., Gandini, R., Sygmund, C., Kittl, R., Haltrich, D., Hallberg, B.M., Ludwig, R., Divne, C. (2015) Structural basis for cellobiose dehydrogenase action during oxidative cellulose degradation. Nat. Commun. 6, 7542. doi: 10.1038/ncomms8542
Tasca, F., Zafar, M.N., Harreither, W., Nöll, G., Ludwig, R., Gorton, L. (2001) A third generation glucose biosensor based on cellobiose dehydrogenase from Corynascus thermophilus and single-walled carbon nanotubes. Analyst 136, 2033-2036. doi: 10.1039/c0an00311e
Wongnate, T. and Chaiyen, P. (2013) The substrate oxidation mechanism of pyranose 2-oxidase and other related enzymes in the glucose-methanol-cholinesuperfamily. FEBS J. 280, 3009-3027. doi: 10.1111/febs.12280
