Isolation and characterization of proteases involved in recombinant protein processing in insect cells
SUPERVISOR: LUKAS MACH
Background.
Baculovirus-based insect cell expression systems are popular platforms for the generation of complex eukaryotic biotherapeutics such as viral surface proteins or multi-subunit complexes used in commercial vaccines (van Oers et al., 2015). Originally, baculovirus-mediated recombinant protein production has been performed mainly in Spodoptera frugiperda cells. Today, Trichoplusia ni cells are recognized as alternative hosts due to their superior capacity to secrete heterologous glycoproteins at high yields (Koczka et al., 2018).
It has been reported that heterologous recombinant proteins can undergo proteolytic processing in insect cells (Bencur et al., 2005; Nika et al., 2017), which can be detrimental for their later use. These cleavage events were initially attributed to v-cath, a papain-like cysteine protease encoded by the baculoviral genome (Hodgson et al., 2013). However, later studies revealed that host proteases such as cathepsin L or the subtilisin-like serine protease furin are also involved in the fragmentation of recombinant proteins produced in insect cells (Johnson and Jiang, 2005; Pagadala et al., 2006).
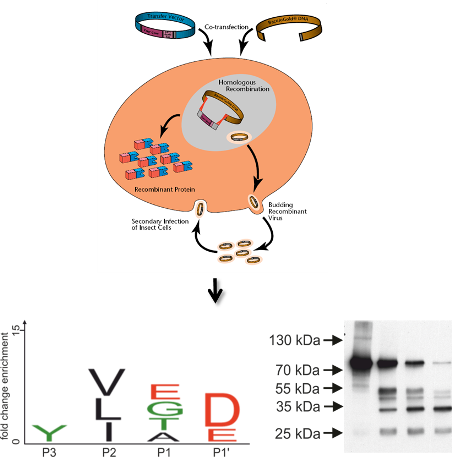
Aims and methods.
We have previously observed that proteolysis of recombinant glycoproteins secreted by insect cells can be reduced by the addition of serine and cysteine protease inhibitors (Bencur et al., 2005). Hence, this project aims at a comprehensive assessment of the serine and cysteine protease activities found in insect cell cultures. Recent progress in the development of activity-based probes for cysteine and serine proteases has allowed the detection of active species of these enzymes in complex samples (van der Hoorn and Kaiser, 2012). Biotin-tagged versions of such functional probes will be used for the selective labelling of cysteine and serine proteases present in extracts and supernatants of S. frugiperda and T. ni cells. After isolation with immobilized streptavidin, the captured proteins will be identified by mass spectrometry-based peptide mapping.
Exploiting the availability of recently obtained transcriptome data (Koczka et al., 2018), the respective cDNAs will be cloned by RT-PCR and then ectopically expressed in insect cells. The recombinant proteases will be purified exploiting exogenously added affinity tags and then analysed for their capacity to degrade particularly vulnerable recombinant proteins (e.g. HER2, influenza hemagglutinin). The cleavage products obtained upon digestion with the recombinant proteases will be characterized by N-terminal sequencing and compared to those generated by incubation with unfractionated cell extracts and culture supernatants. The catalytic properties of the proteases will be investigated in steady-state assays with fluorogenic peptide substrates. Their individual susceptibilities to different synthetic and endogenous protease inhibitors will be established. In addition, the subsite preferences of the enzymes will be determined with proteome-derived peptide libraries and quantitative proteomics after labelling with stable isotopes (Biniossek et al., 2016).
Finally, CRISPR/Cas-based genome editing will be used to generate protease-deficient cell lines which will be tested as hosts for the synthesis of proteolysis-prone heterologous proteins.
Collaborations within this thesis will include ALTMANN (mass spectrometry) and GRABHERR (production of recombinant proteins in insect cells). International collaborations include LENARCIC (enzyme kinetics), SCHILLING (substrate profiling) and VAN DER HOORN (activity-based probes).
Bencur, P., Steinkellner, H., Svoboda, B., Mucha, J., Strasser, R., Kolarich, D., Hann, S., Köllensperger, G., Glössl, J., Altmann, F., Mach, L. (2015) Arabidopsis thaliana beta1,2-xylosyltransferase: an unusual glycosyltransferase with the potential to act at multiple stages of the plant N-glycosylation pathway. Biochem J. 388, 515-525. doi: 10.1042/BJ20042091
Biniossek, M.L., Niemer, M., Maksimchuk, K., Mayer, B., Fuchs, J., Huesgen, P.F., McCafferty, D.G., Turk, B., Fritz, G., Mayer, J., Haecker, G., Mach, L., Schilling, O. (2016) Identification of protease specificity by combining proteome-derived peptide libraries and quantitative proteomics. Mol. Cell. Proteomics 15, 2515-2524. doi: 10.1074/ mcp.O115.056671
Hodgson, J.J., Arif, B.M., Krell, P.J. (2013) Role of interactions between Autographa californica multiple nucleopolyhedrovirus procathepsin and chitinase chitin-binding or active-site domains in viral cathepsin processing. J. Virol. 87, 3471-3483. doi: 10.1128/JVI.01937-12
Johnson, G.D. and Jiang, W. (2005) Characterization of cathepsin L secreted by Sf21 insect cells. Arch. Biochem. Biophys. 444, 7-14. doi: 10.1016/j.abb.2005.09.011
Koczka, K., Peters, P., Ernst, W., Himmelbauer, H., Nika, L., Grabherr, R. (2018) Comparative transcriptome analysis of a Trichoplusia ni cell line reveals distinct host responses to intracellular and secreted protein products expressed by recombinant baculoviruses. J. Biotechnol. 270, 61-69. doi: 10.1016/j.jbiotec.2018.02.001
Nika, L., Wallner, J., Palmberger, D., Koczka, K., Vorauer-Uhl, K., Grabherr, R. (2017) Expression of full-length HER2 protein in Sf9 insect cells and its presentation on the surface of budded virus-like particles. Protein Expr. Purif. 136, 27-38. doi: 10.1016/j.pep.2017.06.005
Pagadala, P.C., Dvorak, L.A., Neet, K.E. (2006) Construction of a mutated pro-nerve growth factor resistant to degradation and suitable for biophysical and cellular utilization. Proc. Natl. Acad. Sci. USA 103, 17939-17943. doi: 10.1073/pnas.0604139103
van der Hoorn, R.A. and Kaiser, M. (2012) Probes for activity-based profiling of plant proteases. Physiol. Plant. 145, 18-27. doi: 10.1111/j.1399-3054.2011.01528.x
van Oers, M.M., Pijlman, G.P., Vlak, J.M. (2015) Thirty years of baculovirus-insect cell protein expression: from dark horse to mainstream technology. J. Gen. Virol. 96, 6-23. doi: 10.1099/vir.0.067108-0
