Simulation of proteins in realistic environments
SUPERVISOR: CHRIS OOSTENBRINK
Background.
The structure and function of complex biomolecules, like proteins, DNA or polysaccharides are studied by molecular simulation which leads to insight into the molecular mechanisms that underlie experimental observations (Karplus and Kuriyan, 2005; van Gunsteren et al., 2006). The quality of molecular simulations should be validated against experimental data (van Gunsteren et al., 2018) and will critically depend on two main aspects: sampling and scoring.
Sampling involves the generation of a sufficiently large amount of molecular structures to describe the conformations that in reality lead to the observed (average) quantities that are measured. These may be obtained from a variety of search techniques, like systematic searches, molecular dynamics or Monte Carlo simulations.
Scoring, on the other hand, is required to evaluate how likely the generated conformations are in terms of energy. The assignment of an energy value to a molecular conformation can again be done in a variety of ways, ranging from advanced quantum mechanical calculations, through force fields to empirical, or knowledge-based scoring functions. Arguably most relevant for biomolecular simulations are atomistic force fields that describe the interactions between the individual atoms using molecular mechanics.
In recent years, we have contributed significantly to the development of the GROMOS force fields for biomolecular simulations (Oostenbrink et al., 2004, Reif et al, 2012). Agreement with experimental data has been explicitly validated on various sets of proteins (Oostenbrink et al., 2005, Reif et al., 2013), and implicitly through the use by the scientific community, leading to approximately 2000 citations of the indicated publications (Web of Science, May 2018). However, the vast majority of all protein simulations reported in the literature, still largely ignores effects of post-translational modifications, co-solvents and crowding on the stability and function of proteins (Feig et al., 2017). We have derived force field parameters for a large set of post-translational modifications (Petrov et al., 2013) and studied the effect of these on the peptide backbones in a preliminary study (Margreitter et al., 2016). Furthermore, we have created a conformational library of carbohydrates relevant in mammalian glycosylation (Turupcu et al., 2017). These works set the stage for a more systematic analysis of effects of post-translational modifications in protein simulations. In addition, effects of e.g. the salt concentration on protein stability have been observed in molecular simulations (e.g. Winger et al., 2007), these largely remain anecdotal and a systematic evaluation of cosolvents in molecular simulations remains unavailable.
Aims and methods.
In this thesis, we will set up a systematic framework to study the effect of cosolvents and post-translational modifications on a wide range of proteins. We have recently established a comprehensive test set of 52 proteins and eight smaller peptides to validate molecular simulations at a larger scale (Setz, 2018).
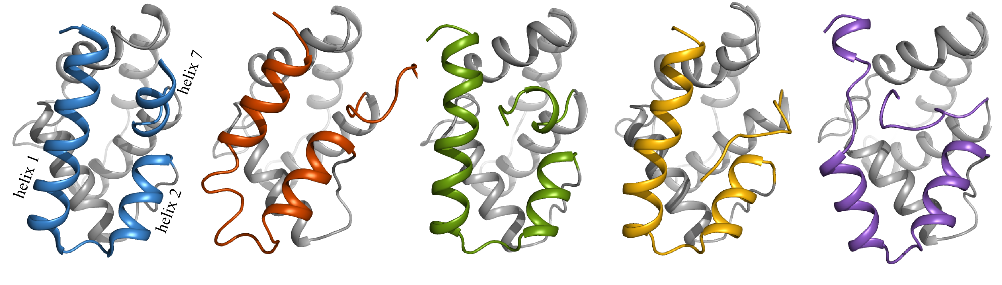
The test set of proteins contains mostly structures derived from X-ray diffraction experiments. These had a resolution smaller or equal to 1.5 Å, no coordinated ions or ligands, did not bind DNA or RNA, and were screened to form monomers in solution and to consist of less than 250 residues. Furthermore, nine structures that were determined by NMR spectroscopy were selected, which were matched against corresponding crystal structures. From simulations with four different force fields statistically significant differences could be observed. At the same time it also became clear that individual proteins may perform very differently between force fields and within replicates, leading to the conclusion that general effects can only be observed by studying significant amounts of biomolecular systems.
Starting from this test set, we will systematically study the effect of different cosolvents on the structure and dynamics of proteins. We will explore if experimentally determined effects can be observed in the timescales of our simulations (nanosecond to microsecond) and study the molecular basis for these effects at a level that is not attainable experimentally. One set of cosolvents will involve the Hofmeister series (Hofmeister, 1888). We will perform molecular dynamics simulations of all proteins in chaotropic and kosmotropic salt solutions, and monitor the effects on structural parameters, like the occurrence of secondary structure elements as well as agreement against experimental observations.
Other cosolvents to be investigated are urea as a denaturant, and DMSO as a solubility enhancer.
Furthermore, we will expand the test set with protein systems that are studied in BioToP to ensure that experimental validation of simulation data can be performed by our collaborators (HALTRICH, JUNGBAUER, KUNERT, LUDWIG, OBINGER). Similarly and using the same simulation machinery, we will collect data on a significant number of post-translationally modified proteins and study the effect of individual modifications on protein structure and dynamics. The effects on e.g. histones (acylation, methylation, phosphorylation), antibodies (glycosylation) and kinases (phosphorylation) will be studied in more detail. By comparing to experimental data, the force-field parameters for the modified amino acids may be validated and possibly refined accordingly.
Collaborations within this thesis include HALTRICH, JUNGBAUER, KUNERT, LUDWIG and OBINGER.
Feig, M., Yu, I., Wang, P.H., Nawrocki, G., Sugita, Y. (2017) Crowding in cellular environments at an atomistic level from computer simulations. J. Phys. Chem. B 121, 8009-8025. doi: 10.1021/acs.jpcb.7b03570
Hofmeister F. (1888) Zur Lehre von der Wirkung der Salze. Zweite Mittheilung. Arch. Exp. Pathol. Pharmacol. 24, 247-260. doi: 10.1007/BF01918191
Karplus, M., Kuriyan, J. (2005) Molecular dynamics and protein functions. Proc. Natl. Acad. Sci. USA 102, 6679-6685. doi: 10.1073/pnas.0408930102
Margreitter, C. and Oostenbrink C. (2016) On the optimization of the protein backbone dihedral angles by means of Hamiltonian reweighting. J. Chem. Inf. Model. 56, 1823-1834. doi: 10.1021/acs.jcim.6b00399
Oostenbrink, C., Villa, A., Mark, A.E., van Gunsteren, W.F. (2004) A biomolecular force field based on the free enthalpy of hydration and solvation: the GROMOS force-field parameter sets 53A5 and 53A6. J. Comput. Chem. 25, 1656-1676. doi: 10.1002/jcc.20090
Oostenbrink, C., Soares, T.A., van der Vegt, N.F.A., van Gunsteren, W.F. (2005) Validation of the 53A6 GROMOS force field. Eur. Biophys. J. 34, 273-284. doi: 10.1007/s00249-004-0448-6
Petrov, D., Margreitter, C., Grandits, M., Oostenbrink, C., Zagrovic, B. (2013) A systematic framework for molecular dynamics simulations of protein posttranslational modifications. PLOS Comput. Biol. 9, e1003154. doi: 10.1371/journal.pcbi.1003154
Reif, M. M., Hünenberger, P. H., Oostenbrink, C. (2012) New interaction parameters for charged amino acid side chains in the GROMOS force field. J. Chem. Theor. Comp. 8, 3705-3723. doi: 10.1021/ct300156h
Reif, M. M., Winger, M., Oostenbrink, C. (2013) Testing of the GROMOS force-field parameter set 54A8: Structural properties of electrolyte solutions, lipid bilayers and proteins. J. Chem. Theor. Comp. 9, 1247-1264. doi: 10.1021/ct300874c
Setz, M. (2018) Molecular Dynamics Simulations of Biomolecules: From Validation to Application. PhD thesis, University of Natural Resources and Life Sciences, Vienna.
Turupcu, A. and Oostenbrink, C. (2017) Modeling of Oligosaccharides within Glycoproteins from Free-Energy Landscapes. J. Chem. Inf. Model. 57, 2222-2236. doi: 10.1021/acs.jcim.7b00351
van Gunsteren, W.F., Bakowies, D., Baron, R., Chandrasekhar, I., Christen, M., Daura, X., Gee, P., Geerke, D., Glättli, A., Hünenberger, P.H., Kastenholz, M. A., Oostenbrink, C., Schenk, M., Trzesniak, D., van der Vegt, N. F. A., Yu, H.B. (2006) Biomolecular modeling: goals, problems, perspectives. Angew. Chem. Int. Ed. 45, 4064-4092. doi: 10.1002/ anie.200502655
van Gunsteren, W.F., Daura, X., Hansen, N., Mark, A.E., Oostenbrink, C., Riniker, S., Smith L.J. (2018) Validation of molecular simulation: an overview of issues. Angew. Chem. Intl. Ed. 57, 884-902. doi: 10.1002/anie.201702945
Winger, M. Yu, H.B., Redfield, C., van Gunsteren, W.F. (2007) Molecular dynamics simulation of human interleukin-4: comparison with NMR data and effect of pH, counterions and force field on tertiary structure stability. Mol. Sim. 33, 1143-1154. doi: 10.1080/08927020701613623
