Redox communication between cytosol and ER
JOINT-SUPERVISORS: Brigitte GASSER and Stephan HANN
Background.
In eukaryotes, formation of disulfide bonds in secretory proteins is attained through the oxidative protein folding machinery in the endoplasmic reticulum (ER), using glutathione as redox buffer. Glutathione itself is converted from its oxidized form (GSSG) to its reduced form (GSH) by dedicated enzymes localized in the cytosol or mitochondria, which require NADPH as cofactor. We have previously demonstrated a strong impact of secretory protein production on cellular redox state and vice versa. Modulation of the oxidation state of ER in the yeast Pichia pastoris (syn Komagataella spp) enhanced protein folding and secretion, as did re-balancing of the cytosolic redox state (Delic et al. 2012; Delic et al. 2014).
Based on the fact that we observed a strong redox communication between the cytosol and the ER, we hypothesize that there must be an ER membrane transporter for the important redox factor glutathione. This is also underlined by the fact that glutathione can only be synthesized in the cytosol, but is also present in the organelles such as ER and mitochondria (Bachhawat et al. 2013). As each organelle seems to able to regulate the redox state of its glutathione pools, and needs to be able to transport both GSH and GSSG across its membranes, the elucidation of redox communication between compartments is of high interest (Elbaz-Alon et al. 2014). Despite being proposed for several years, so far no candidate transporter for the ER has been reported, and it is not clear whether the oxidized or reduced forms of glutathione are the transported molecules. One limit in this respect is that accurate determination of GSH and GSSG concentration in biological systems is extremely challenging as several sources of error have to be considered and assessed. Evidently, these challenges are even increasing when it comes to the quantification of the compounds in individual intracellular organelles such as the ER, as several centrifugation and pre-concentration steps need to be performed during their isolation. As a consequence, both the absolute concentration of GSH and GSSG as well as their ratio may be changed, thus means to monitor and quantify GSH oxidation during both cell lysis/extraction and organelle isolation need to be established and implemented. Some years ago, we have developed and successfully applied a method, which utilizes a stable isotope labeled internal standard to correct some of these effects when quantifying GSH and GSSG in whole cell extracts (Haberhauer-Troyer et al. 2013).
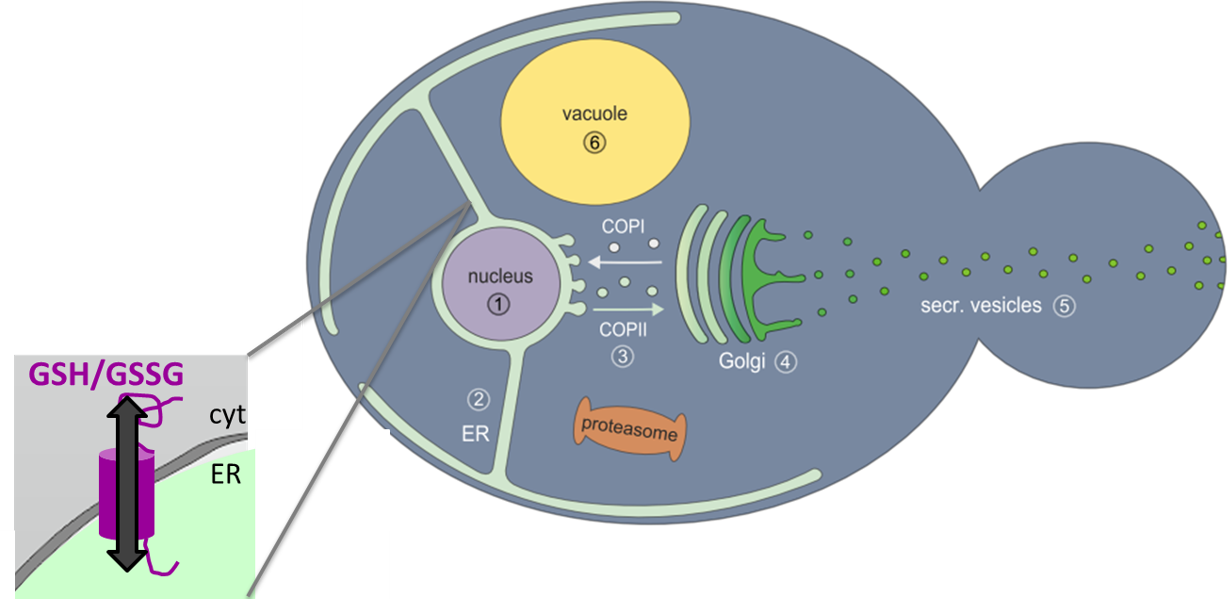
Recently, it has been published that GSH import into the ER is occurring by facilitated diffusion via the translocon pore in S. cerevisiae when artificially enriched glutathione pools and consequent ER stress were applied (Ponsero et al. 2017), however, no evidence for ER glutathione export via this pathway was provided, and transport was not studied under normal conditions. Furthermore, given the fact that other organelles only allow directed import and export of glutathione via dedicated transporters (Elbaz-Alon et al. 2014), the additional presence of a dedicated ER glutathione transporter seems plausible. Such a transporter needs to be localized in the ER membrane and contain glutathione binding sites (Bachhawat et al. 2013). Based on bioinformatics analyses, we have identified one potential candidate named Emp1, which is localized in the ER membrane and carries a potential glutathione transport motif. In preliminary experiments Emp1 was shown to be essential for viability and to influence ER glutathione levels, leading to a more oxidizing ER environment upon overexpression.
Research Questions:
Is the identified ER membrane protein Emp1 acting as glutathione transporter between ER and cytosol?
Which is its mode of action (transport of GSH or GSSG) and which cofactors and interaction partners are needed therefore?
Which bioanalytical strategies are fit-for-purpose regarding the isolation of cellular compartments and redox metabolites with high stability and yield?
Can stable isotope labelling during cell cultivation in combination with mass spectrometric analysis advance the accuracy and robustness of intracellular redox state assessment?
Objectives:
Characterize Emp1 function and interaction partners
Establish analytical methods suitable to accurately quantify redox metabolites such as GSH and GSSG in organelle isolations
Employ knowledge for increasing recombinant protein secretion
Aims and methods.
The aims of this project are to characterize the putative transporter responsible for ER-cytosol glutathione exchange in more detail with the aim to elucidate its function and interaction partners, and to establish suitable fit-for-purpose analytical methods to accurately quantify GSH and GSSG in organelle preparations. Therefore, strains bearing different mutations in the predicted glutathione binding sites of the putative transporter and tagged variants of the transporter will be constructed and analyzed. The tagged version of the transporter will be used to determine its orientation within the ER membrane (by immunofluorescence microscopy) and its interaction partners (by co-immunoprecipitation and LC-MS/MS, ALTMANN). Interaction partners will provide one step towards elucidating the mode of action: (i) It is assumed that the energy required for this transportation step (e.g. in form of ATP or GTP) is provided by an interaction partner. (ii) If GSSG is the transported cargo, its formation is assumed to be close to the transporter by a dedicated enzyme (potential candidates include Ero1 on the ER side or Fmo1 on the cytosolic side, Suh et al. 1999)
To determine whether GSH or GSSG are transported, ER (microsomes) of the mutant strains will be isolated (according to the protocol by Klug et al. 2014) and reduced and oxidized glutathione will be accurately quantified by advancing the method introduced by Haberhauer-Troyer et al. in 2013 for whole cell extracts of yeast, which utilizes a stable isotope labeled internal standard. As a drawback the addition of stable isotope labeled internal standards is, in general, performed during the extraction of the target compounds and cannot correct extraction efficiency, but only errors occurring due to compound degradation or losses. Therefore the analytical research in this project focuses on the development and implementation of advanced stable isotope labeling strategies. For the first time stable isotope labelling during cell cultivation will be applied in the context of sulfur metabolomics. In a differential approach, the addition of either 34S-labeled sulfates (or 13C-labeled glucose) will be performed with one strain (possessing e.g. a mutant of the transporter), whereas the second strain (containing e.g. the wild type transporter) will be cultivated on natural substrates. Before the conduction of the organelle-specific isolation methods the cells of the two cultivations will be combined. As a consequence mass spectrometric analysis can be performed by LC-MS in the combined strains by simultaneous monitoring of GnatSH, G34SH, GnatSnatSG and G34S34SG – it is noteworthy that this is a so-called ration method enabling the delivery of the accurate ratio of GSH/GSSG in the organelles of two selected strains. Formation of other (mixed) species such as GnatS34SG will be carefully evaluated and the ratios will be corrected accordingly. In addition, our competence and equipment enable us to determine 32S/34S with highest sensitivity and accuracy using LC-ICPMS.

However, as this method does not offer the high selectivity of molecular mass spectrometry, the selective chromatographic separation of GSH and GSSC from other sulfur-containing molecule is a prerequisite and needs careful evaluation. Furthermore, redox-sensitive GFP variants will be applied to study organelle-specific redox conditions in vivo (Delic et al. 2010) in the mutant strains. Additionally, marker genes for ER and cytosolic stress will be analysed by RT-qPCR in the mutant strains and their impact on recombinant protein secretion will be assessed. Together, these approaches will enable the understanding of glutathione transport between the cytosol and the ER.
Collaborations and time plan:
Generation of P. pastoris strains with alanine scanning versions of the putative transporter and analysis with in vivo fluorescent redox biosensors (6 months in the 1st year), generation of tagged versions for proteomics and microscopy as well as generation of knockout or overexpression mutants of interaction partners and all cultivations (4-6 months each in years 2 and 3) will be performed in the group of Brigitte GASSER. Establishment of the differential isotope labelling and subsequent measurement of redox active metabolites (especially GSH and GSSG) during and after microsome isolation of transporting and non-transporting strains will take place in the group of Stephan HANN (9-12 months in the 1st/2nd year plus 6 months in the last year). Further collaborations within this thesis will include the group of Friedrich ALTMANN for proteomics to identify interaction partners (measurements to be performed by BioToP faculty; for methods see: Pfeffer et al. 2012, Klug et al. 2014, Burgard et al. 2019).
A stay abroad of 6 months is anticipated, depending on the PhD student´s background this might be e.g. in the groups of Coral BARBAS (University Foundation San Pablo CEU, Madrid) or Lloyd RUDDOCK (University of Oulu, Finnland).
Bachhawat, A.K., Thakur, A., Kaur, J., Zulkifli, M. (2013) Glutathione tranporters. Biochim Biophys Acta. 1830, 3154-64. doi: 10.1016/j.bbagen.2012.11.018.
Burgard J., Valli M., Grünwald-Gruber C., Zanghellini J., Altmann F., Mattanovich D., Gasser B. (2019) The secretome of Pichia pastoris in fed-batch cultivations is largely independent of the carbon source but changes quantitatively over cultivation time. Microb Biotechnol. Accepted.
Delic, M., Mattanovich, D., Gasser, B. (2010) Monitoring intracellular redox conditions in the endoplasmic reticulum of living yeasts. FEMS Microbiol Lett. 306, 61-6. doi: 10.1111/j.1574-6968.2010.01935.x.
Delic, M., Rebnegger, C., Wanka, F., Puxbaum, V., Haberhauer-Troyer, C., Hann, S., Koellensperger G., Mattanovich D., Gasser B. (2012). Oxidative protein folding and unfolded protein response elicit different redox regulation in ER and cytosol of yeast. Free Radic Biol Med. 52(9): 2000-12. doi: 10.1016/j.freeradbiomed.2012.02.048.
Delic, M., Graf, A.B., Koellensperger, G., Haberhauer-Troyer, C., Hann, S., Mattanovich, D., Gasser, B. (2014) Overexpression of the transcription factor Yap1 modifies intracellular redox conditions and enhances recombinant protein secretion. Microb. Cell 1, 376-386. doi: 10.15698/mic2014.11.173.
Elbaz-Alon, Y., Morgan, B., Clancy, A., Amoako, T.N., Zalckvar, E., Dick, T.P., Schwappach, B., Schuldiner, M. (2014) The yeast oligopeptide transporter Opt2 is localized to peroxisomes and affects glutathione redox homeostasis. FEMS Yeast Res. 14, 1055-67. doi: 10.1111/1567-1364.12196.
Haberhauer-Troyer, C., Delic, M., Gasser, B., Mattanovich, D., Hann, S., Koellensperger, G. (2013) Accurate quantification of the redox-sensitive GSH/GSSG ratios in the yeast Pichia pastoris by HILIC-MS/MS. Anal. Bioanal. Chem. 405, 2031-2039. doi: 10.1007/s00216-012-6620-4.
Klug, L., Tarazona, P., Gruber, C., Grillitsch, K., Gasser, B., Trötzmüller, M., Köfeler, H., Leitner, E., Feussner, I., Mattanovich, D., Altmann, F., Daum, G. (2014) The lipidome and proteome of microsomes from the methylotrophic yeast Pichia pastoris. Biochim Biophys Acta. 1841, 215-26. doi:pii: S1388-1981(13)00247-3. 10.1016/j.bbalip.2013.11.005.
Pfeffer M., Maurer M., Stadlmann J., Grass J., Delic M., Altmann F., Mattanovich D. (2012) Intracellular interactome of secreted antibody Fab fragment in Pichia pastoris reveals its routes of secretion and degradation Appl Microbiol Biotechnol. 93(6), 2503-12. doi: 10.1007/s00253-012-3933-3.
Ponsero, A.J., Igbaria, A., Darch, M.A., Miled, S., Outten, C.E., Winther, J.R., Palais, G., D'Autréaux, B., Delaunay-Moisan, A., Toledano, M.B. (2017) Endoplasmic reticulum transport of glutathione transport by Sec61 is regulated by Ero1 and Bip. Mol Cell. 67, 962-973.e5. doi: 10.1016/j.molcel.2017.08.012.
Suh, J.K., Poulsen, L.L., Ziegler, D.M., Robertus, J.D. (1999) Yeast flavin-containing monooxygenase generates oxidizing equivalents that control protein folding in the endoplasmic reticulum. Proc Natl Acad Sci U S A. 96, 2687-91.
