Enzymatic properties and evolutionary origin of the cancer-associated protease cathepsin O
SUPERVISORS: Dietmar HALTRICH and Lukas MACH
Background.
Lysosomal cysteine proteinases (cysteine cathepsins) play important roles in a wide range of physiological processes including antigen presentation (van Kasteren and Overkleeft, 2014), bone resorption (Novinec and Lenarcic, 2013) and programmed cell death (Repnik et al., 2012). Furthermore, cysteine cathepsins are of substantial pathological relevance in disease states such as cancer (Olson and Joyce, 2015). Cysteine cathepsins belong to the so-called papain superfamily of cysteine proteinases, which includes eleven human representatives (Ketterer et al., 2017). The catalytic and structural properties of most human cathepsins are already well characterized (Turk et al., 2012). Cathepsin O is the only human cysteine cathepsin that has so far received little attention (Reif et al., 2012). However, a recent study has revealed that cathepsin O seems to determine the drug sensitivity of human breast cancers (Cairns et al., 2017).
Aims and methods.
This project aims at a comprehensive characterization of the enzymatic properties and evolutionary origin of cathepsin O. This protease is ubiquitously expressed in human tissues (Velasco et al., 1994). Cathepsin O shares all major sequence features of cathepsin L-like cysteine proteinases, including the conserved active-site residues Cys132, His269 and Asn289 forming the catalytic triad. However, the propeptide of cathepsin O is considerably shorter than in the case of cathepsin L and other related enzymes (Reif et al., 2012). This could compromise the molecular functions of the propeptide, which is generally considered as necessary for folding and activation of cysteine cathepsins (Müntener et al., 2005). Interestingly, the chromosomal localization and exon/intron arrangement of the human cathepsin O gene is unique, indicating evolutionary distance from the other cysteine cathepsin genes (Velasco et al., 1998). An ancient origin of cathepsin O is also supported by its occurrence in primitive organisms such as choanoflagellates and sponges (Riesgo et al., 2015). However, some characteristic sequence motifs of human cathepsin O are not entirely conserved in its evolutionary precursors. This could reflect a functional adaptation of the enzymatic properties of cathepsin O over time.
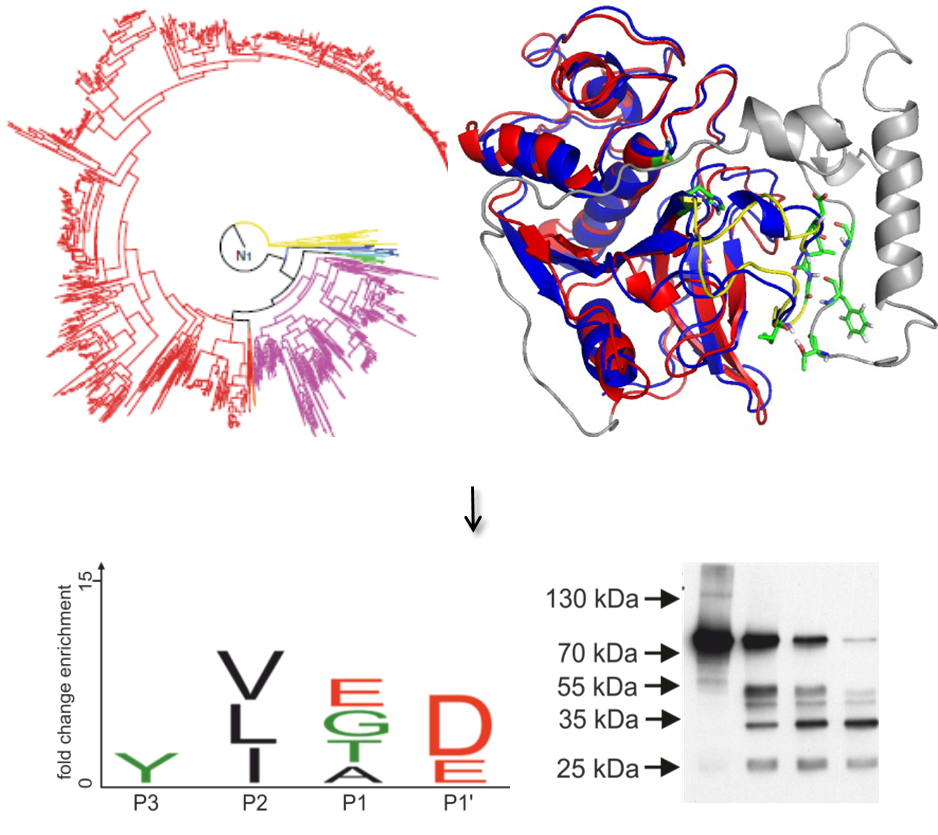
The early appearance of cathepsin O during animal evolution provides sufficient sequence space to reconstruct its evolutionary ancestor as recently described for other ancient enzymes (Gumulya et al., 2018). The first goal of this project is therefore the functional expression of natural and reconstructed cathepsin O precursors in Escherichia coli. Purified recombinant human and ancestral procathepsin O will then be tested for self-removal of their propeptides as typical for papain-like cysteine proteinases (Niemer et al., 2016). The catalytic properties of the enzymes will be investigated in steady-state assays with fluorogenic peptide substrates (Paireder et al., 2017). Their individual susceptibilities to different synthetic and endogenous protease inhibitors will be established. In addition, the subsite preferences of the proteases will be determined with proteome-derived peptide libraries and quantitative proteomics after labelling with stable isotopes (Biniossek et al., 2016). This information will then be exploited for the design of optimized internally quenched fluorescence substrates and activity-based probes for the selective detection of cathepsin O in complex samples such as tumour extracts. Molecular modelling based on the closely related procathepsin L structure (Reif et al., 2012) will be used to identify regions of human cathepsin O suitable for saturation mutagenesis to further characterize the functional relevance of the unique architecture of the cathepsin O propeptide. Efforts will be undertaken to determine the three-dimensional structure of human and ancestral cathepsin O by X-ray crystallography. Collectively, these studies shall provide unprecedented insights into the catalytic significance of the unique molecular features of cathepsin O and thus set the stage for detailed studies on the functions of this enzyme in human cancers.
This project will be jointly supervised by MACH (protein expression, protease characterization) and HALTRICH (ancestor reconstruction, protein crystallization). Additional internal collaboration partners are PETERBAUER (saturation mutagenesis) and OOSTENBRINK (molecular modelling). International collaborations include NOVINEC (structure determination) and SCHILLING (substrate profiling).
Biniossek ML, Niemer M, Maksimchuk K, Mayer B, Fuchs J, Huesgen PF, McCafferty DG, Turk B, Fritz G, Mayer J, Haecker G, Mach L, Schilling O. Identification of protease specificity by combining proteome-derived peptide libraries and quantitative proteomics. Mol Cell Proteomics. 2016;15:2515-24
Cairns J, Ingle JN, Wickerham LD, Weinshilboum R, Liu M, Wang L. SNPs near the cysteine proteinase cathepsin O gene (CTSO) determine tamoxifen sensitivity in ERα-positive breast cancer through regulation of BRCA1. PLoS Genet. 2017;13:e1007031.
Gumulya Y, et al. Engineering highly functional thermostable proteins using ancestral sequence reconstruction. Nature Catalysis. 2018;1: 878-88.
Ketterer S, Gomez-Auli A, Hillebrand LE, Petrera A, Ketscher A, Reinheckel T. Inherited diseases caused by mutations in cathepsin protease genes. FEBS J. 2017;284:1437-54
Müntener K, Willimann A, Zwicky R, Svoboda B, Mach L, Baici A. Folding competence of N-terminally truncated forms of human procathepsin B. J Biol Chem. 2005;280:11973-80
Niemer M, Mehofer U, Verdianz M, Porodko A, Schähs P, Kracher D, Lenarcic B, Novinec M, Mach L. Nicotiana benthamiana cathepsin B displays distinct enzymatic features which differ from its human relative and aleurain-like protease. Biochimie. 2016;122:119-25
Novinec M, Lenarcic B. Cathepsin K: a unique collagenolytic cysteine peptidase. Biol Chem. 2013;394:1163-79
Olson OC, Joyce JA. Cysteine cathepsin proteases: regulators of cancer progression and therapeutic response. Nat Rev Cancer. 2015;15:712-29.
Paireder M, Tholen S, Porodko A, Biniossek ML, Mayer B, Novinec M, Schilling O, Mach L. The papain-like cysteine proteinases NbCysP6 and NbCysP7 are highly processive enzymes with substrate specificities complementary to Nicotiana benthamiana cathepsin B. Biochim Biophys Acta Proteins Proteom. 2017;1865:444-452
Reif MM, Mach L, Oostenbrink C. Molecular insight into propeptide-protein interactions in cathepsins L and O. Biochemistry. 2012;51:8636-53.
Repnik U, Stoka V, Turk V, Turk B. Lysosomes and lysosomal cathepsins in cell death. Biochim Biophys Acta. 2012;1824:22-33.
Riesgo A, Maldonado M, López-Legentil S, Giribet G. A proposal for the evolution of cathepsin and silicatein in sponges. J Mol Evol. 2015;80:278-91
Santamaría I, Pendás AM, Velasco G, López-Otín C. Genomic structure and chromosomal localization of the human cathepsin O gene (CTSO). Genomics. 1998;53:231-4.
Turk V, Stoka V, Vasiljeva O, Renko M, Sun T, Turk B, Turk D. Cysteine cathepsins: from structure, function and regulation to new frontiers. Biochim Biophys Acta. 2012;1824:68-88.
van Kasteren SI, Overkleeft HS. Endo-lysosomal proteases in antigen presentation. Curr Opin Chem Biol. 2014;23:8-15.
Velasco G, Ferrando AA, Puente XS, Sánchez LM, López-Otín C. Human cathepsin O. Molecular cloning from a breast carcinoma, production of the active enzyme in Escherichia coli, and expression analysis in human tissues. J Biol Chem. 1994;269:27136-42.
