The cancer immunotherapy antigen CD19 and its molecular interaction network
SUPERVISORS: Renate KUNERT and Christian OBINGER
Background.
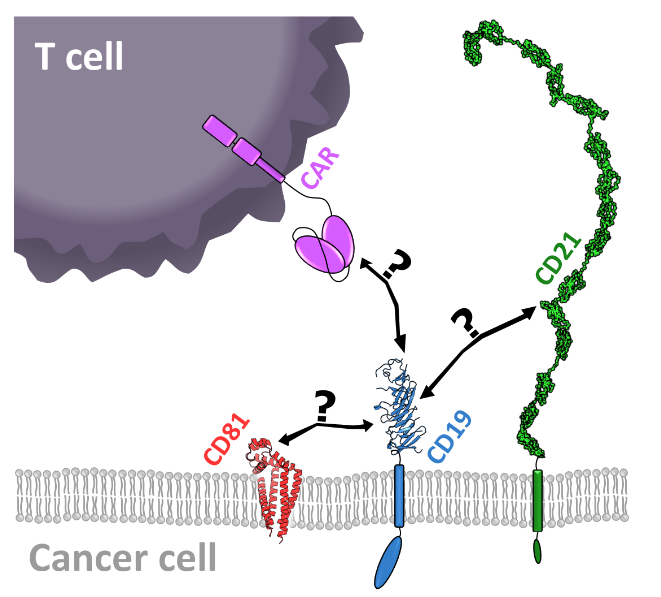
CD19 is a transmembrane protein that is found on the surface of B cells. Since its expression is restricted to B cells, CD19 is perfectly suited as an antigen for immunotherapies targeting B cell-derived tumors, such as acute lymphoblastic leukaemia (ALL) or B cell lymphomas. Different types of CD19-targeting therapeutics have been approved, including chimeric antigen receptor (CAR) T cells and the bispecific antibody blinatumomab (Feins et al., 2019). Despite its central role in cancer immunotherapies, surprisingly little is known about its interaction network within the B cell coreceptor complex consisting of CD19, CD21 and CD81 (see Figure).
This lack of understanding may be partly explained by the fact that the extracellular domain of CD19 (CD19-ECD) is considered as being a “difficult-to-express” (DTE) protein. Apart from being expressed at very low titers, the CD19-ECD misfolds and aggregates, which precludes detailed interaction studies. Importantly, we have recently engineered CD19-ECD mutants which can be expressed at high yields and fold into the native CD19-structure, which provides the basis for CD19-based interaction studies in this thesis.
Aims and methods.
The goal of this project is to generate a detailed understanding of how the different components of the B cell coreceptor complex interact with the CD19 molecule. Moreover, it will be investigated how those interactions influence the potency of CD19-targeting CAR T cell immunotherapies, e.g., by competitively binding to the same epitope (i.e., binding sites) on CD19 or by inducing conformational changes in this antigen. Thus, elucidating the interaction between those membrane proteins might contribute to the development of improved tumour treatment strategies, e.g., by blocking or targeting newly defined active regions on the protein.
As described above, we have recently established the expression of improved CD19-ECD mutants in HEK cells. Apart from those improved CD19-ECD variants, the extracellular domains of the interaction partners CD81 and CD21 will also be expressed in mammalian host cell lines (e.g., CHO or HEK cells), which are established in our laboratory (Hennicke et al., 2019; Mader et al., 2013) . The purity and folding properties of all proteins will be analysed by SEC-MALS and DSC (core facility BmCA). Based on those solubly expressed and purified proteins a broad spectrum of biochemical interaction studies will be conducted. First, the epitopes of CD81, CD21 and the clinically approved CARs will be defined at high resolution. For that purpose, we will make use of a randomly mutated CD19-ECD-library that is expressed on the surface of yeast cells, i.e., a yeast display library. This library will be selected for non-binding to those different ligands (CD81, CD21 and the recognition domain of the anti-CD19 CAR), which will yield a set of mutations for each interaction partner, which are located in the respective epitope. Of note, we have successfully used a similar yeast display-based approach in the past for epitope mapping on IgG1-Fc (Traxlmayr et al., 2012) . Ultimately, this first part of the project will identify the interaction sites of those ligands on the CD19-ECD molecule. To further extend the knowledge on the CD19 interaction web, the affinities of the CD19-ECD to its ligands will be determined by titration on CD21- and CD81-positive cell lines, respectively (cooperation with GRILLARI). This affinity analysis will be complemented by SPR analysis (core facility BmCA). Next, to obtain information on the CD19-CD81 interaction at atomic resolution, the extracellular domains of those two proteins will be co-crystallized to resolve the structure of the assembled complex (cooperation with DJINOVIC-CARUGO), similar to our previous studies on antibody-antigen complexes (Lobner et al., 2017a; Lobner et al., 2017b) . This structure will give valuable insight on the orientation of the CD19-ECD on the B cell surface (caused by tethering to the CD81 transmembrane protein). Moreover, it will be interesting to compare the CD19-CD81 complex structure with that of unboundCD19 (Teplyakov et al., 2018), which will show whether the interaction with its folding chaperone CD81 induces a conformational change. Such a conformational switch would of course have direct implications on the recognition by CAR T cells. Finally, depending on the outcome of those experiments, the influence of various parameters (e.g., CD81 expression density, addition of soluble CD81-ECD or CD21 expression density) on the efficacy of CAR T cells will be analysed (cooperation with LEHNER). This will yield highly valuable information, which might explain some of the CAR T cell resistances observed in cancer patients after therapy.
Collaborations within this thesis will include GRILLARI (CD81-positive cell lines). National collaborations will include LEHNER (CAR T cells) and DJINOVIC-CARUGO (X-ray crystallography).
Feins, S., Kong, W., Williams, E.F., Milone, M.C., and Fraietta, J.A. (2019). An introduction to chimeric antigen receptor (CAR) T-cell immunotherapy for human cancer. American journal of hematology.
Hennicke, J., Reinhart, D., Altmann, F., and Kunert, R. (2019). Impact of temperature and pH on recombinant human IgM quality attributes and productivity. New biotechnology 50, 20-26.
Lobner, E., Humm, A.S., Goritzer, K., Mlynek, G., Puchinger, M.G., Hasenhindl, C., Ruker, F., Traxlmayr, M.W., Djinovic-Carugo, K., and Obinger, C. (2017a). Fcab-HER2 Interaction: a Menage a Trois. Lessons from X-Ray and Solution Studies. Structure 25, 878-889 e875.
Lobner, E., Humm, A.S., Mlynek, G., Kubinger, K., Kitzmuller, M., Traxlmayr, M.W., Djinovic-Carugo, K., and Obinger, C. (2017b). Two-faced Fcab prevents polymerization with VEGF and reveals thermodynamics and the 2.15 A crystal structure of the complex. mAbs 9, 1088-1104.
Mader, A., Prewein, B., Zboray, K., Casanova, E., and Kunert, R. (2013). Exploration of BAC versus plasmid expression vectors in recombinant CHO cells. Applied microbiology and biotechnology 97, 4049-4054.
Teplyakov, A., Obmolova, G., Luo, J., and Gilliland, G.L. (2018). Crystal structure of B-cell co-receptor CD19 in complex with antibody B43 reveals an unexpected fold. Proteins 86, 495-500.
Traxlmayr, M.W., Hasenhindl, C., Hackl, M., Stadlmayr, G., Rybka, J.D., Borth, N., Grillari, J., Ruker, F., and Obinger, C. (2012). Construction of a stability landscape of the CH3 domain of human IgG1 by combining directed evolution with high throughput sequencing. Journal of molecular biology 423, 397-412.
