Mechanistic synergism between bacterial pyranose oxidase and peroxidase in lignin depolymerization
SUPERVISORS: CLEMENS PETERBAUER and CHRIS OOSTENBRINK
Background:
The auxiliary activity family 3 (AA3; CAZy, http://www.cazy.org/) comprises flavin-dependent oxidoreductases such as cellobiose dehydrogenase, aryl alcohol oxidase, glucose oxidase, glucose dehydrogenase, and pyranose oxidase (Cavener et al., 1992; Sützl et al., 2018). Their reaction mechanism involves a hydride transfer from the substrate resulting in reduction of the flavin cofactor (FAD) to FADH2 (reductive half-reaction). FADH2 is re-oxidized (oxidative half-reaction) by an electron acceptor (Wongnate et al., 2013), which can be molecular oxygen (in oxidases; reduced to hydrogen peroxide), as well as a range of (substituted) quinones, (complexed) metal ions, or even various radicals (Leitner et al., 2001), in dehydrogenases. In wood-degrading basidiomycetes, oxidases are considered to provide hydrogen peroxide to activate lignin-modifying peroxidases (formation of compound I). Dehydrogenases are implicated in maintaining a quinone/hydroquinone redox cycle as well as in the provision of reduced metals for radical-based depolymerization reactions (Kracher et al., 2016). Pyranose oxidase (subfamily AA3_4) and other AA3 oxidases often show a pronounced dehydrogenase activity. Recently, POx from the white-rot basidiomycete Irpex lacteus was shown to reduce quinoid intermediates produced by laccase or peroxidases in vitro and prevent their (re)polymerization (Ai et al., 2014; Wang et al. 2019).
Research on lignin depolymerization has focused on fungal systems thus far (Cragg et al., 2015; Pollegioni et al., 2015; Janusz et al., 2017), however, lignin oxidation was observed in a number of soil bacteria, mostly from the taxonomic groups of Actinobacteria, α-Proteobacteria and γ-Proteobacteria (Cragg et al., 2015). Recent studies implicate dye-decolorizing peroxidases (DyP) to be key enzymes in bacterial lignin depolymerization (Brown et al., 2012; Brown and Chang, 2014). While biochemical data on these enzymes are limited, certain bacterial DyP were shown to possess an activity comparable to fungal DyP and manganese peroxidases (Min et al., 2015).
We searched for putative AA3-family enzymes in bacterial genomes and found only enzymes with similarity to fungal POx, with most genes occuring in Actinobacteria and Proteobacteria. We subsequently expressed a novel pyranose oxidase from Kitasatospora aureofaciens (formerly Streptomyces aureofaciens; KaPOx) and found substantial similarities with ToPOx from Trametes ochracea, the best-characterized fungal POx. The enzyme was able to utilize similar two- and one-electron acceptors; additionally, complexed Mn(III) was found to be a suitable one-electron acceptor. We observed redox cycling of model compounds between KaPOx and a manganese peroxidase: MnP catalyzed the oxidation of 2,6-dimethoxyphenol (DMP), guaiacol, acetosyringone and sinapic acid to their respective quinoids. Addition of KaPOx to the mixture caused a full or partial reversion of the reaction. Conversely, reduction of 1,4-benzoquinone (1,4-BQ) by KaPOx could be reverted through partial re-oxidation by MnP (Herzog, Sützl et al., manuscript in revision).
Aims:
Based on these results we propose that certain Actinobacteria employ DyP (six putative genes are found in the K. aureofaciens genome) and POx for lignin oxidation. The role for POx in this process would be a double one: using available monosaccharides as substrate, POx can supply H2O2 to activate peroxidases for lignin decomposition at the onset of lignin depolymerization. The peroxidase reaction products, aromatic lignin compounds (radicals) and highly reactive Mn(III) ions, can be reduced by the dehydrogenase activity and thus prevent their repolymerization as well as damage to cellular components.
The major questions are:
- Are bacterial DyP capable of producing the radical or metal mediators from available substrates upon activation with hydrogen peroxide to effect depolymerization of lignin?
- How are bacterial DyP differentiated from fungal DyP (and fungal LiP and MnP) in terms of reactivity?
- Are the produced aromatic radicals or metal ions suitable electron acceptors for the oxidative half-reaction of bacterial POx?
- How do the proposed electron acceptors (peroxidase reaction products) interact with POx?
Methods:
We plan to express the candidate genes for DyP and POx from the model organism K. aureofaciens in suitable expression systems and characterize the resulting enzymes with respect to their substrate specificity and product range. We will use molecular modeling to construct homology models of KaPOx and subsequently dock peroxidase reaction products into the active site. Diffusion of molecular oxygen within the protein can be studied using molecular dynamics simulations (Baron et al., 2009). KaPOx is the first POx where Mn(III) ions, reaction products of Mn-peroxidases and key mediators in white-rot lignin depolymerization, were shown to be a suitable electron acceptor (Herzog, Sützl et al., manuscript in revision). We will thus include Mn(III) ions, alone and complexed by organic acids such as oxalate, in these modeling experiments. From these models, we will identify key amino acid residues or motifs in KaPOx that are involved in the use of molecular oxygen and alternative electron acceptors. The results will be used as guidelines for enzyme mutagenesis/engineering, in order to determine the molecular basis of how the enzyme “switches” between oxidase and dehydrogenase activity.
Once a putative DyP-type peroxidase is confirmed and characterized experimentally, we will also create in silico models for this enzyme and attempt to elucidate the reaction mechanism and differences in substrate and product range compared to established fungal lignin and manganese peroxidases. This will allow us to determine whether such lignin-modifying bacteria are capable of facilitating oxidative attack on lignin by themselves, or whether they are reliant on the activity of fungi that occupy the same habitat.
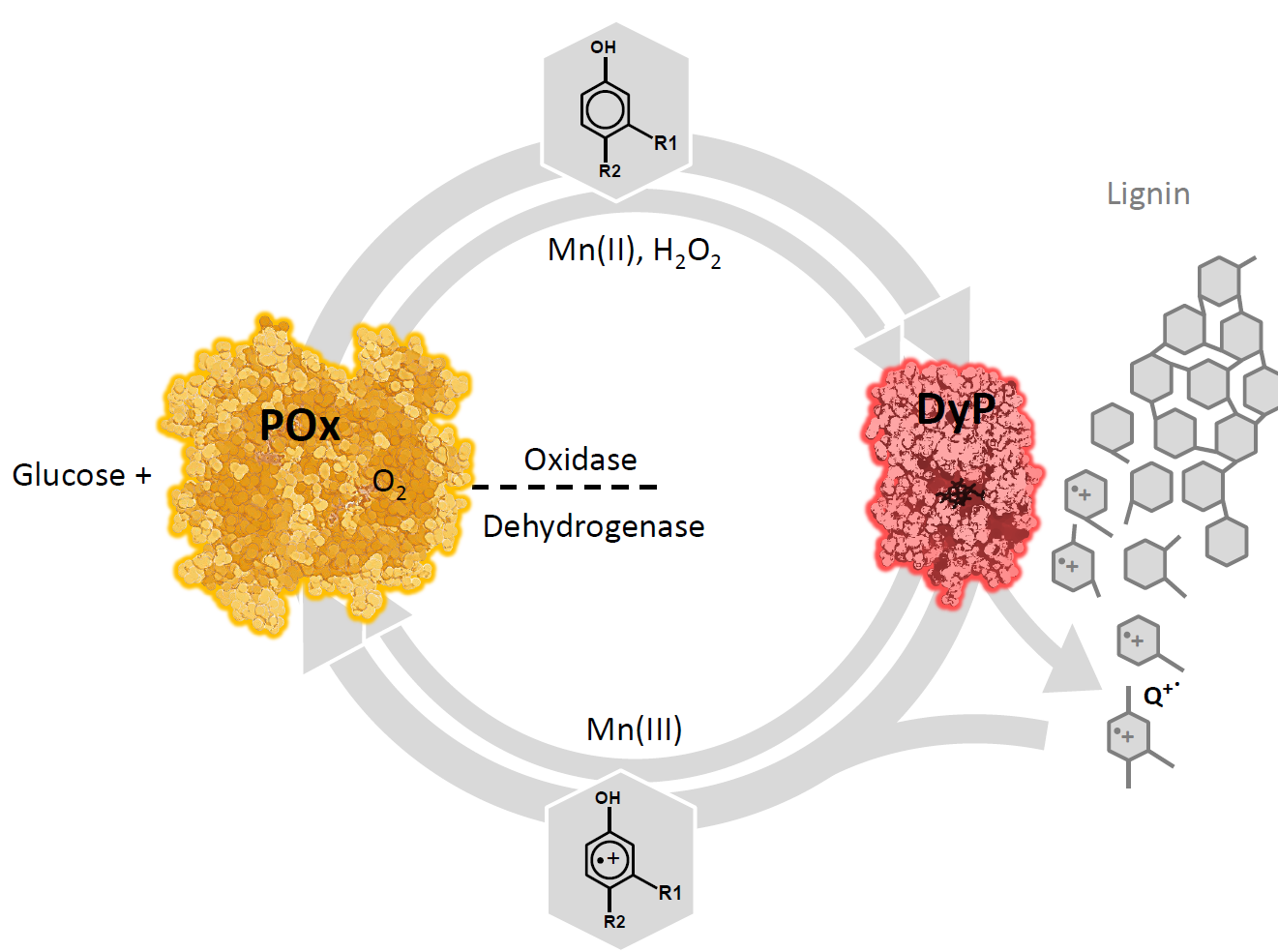
Proposed role of POx as a redox partner of peroxidases: POx oxidizes monosaccharides (D-glucose and others) and supplies H2O2 (oxidase activity) for peroxidases (DyP), which produce aromatic radicals in lignin. These radicals are (partially) reduced by POx (dehydrogenase activity, which prevents re-polymerization and cellular damage, and shifts the balance towards degradation. For some peroxidases, Mn(III) is a mediator in this process. It can concomitantly be reduced, thus detoxified and recycled by POx.
Collaborations: This thesis is envisioned as a joint project between PETERBAUER (protein engineering and characterization) and OOSTENBRINK (molecular modelling and simulations) and Collaborations within BioToP are foreseen with OBINGER (protein characterization).
REFERENCES:
Ai M-Q, Wang F-F, Zhang Y-Z, Huang F. 2014. Purification of pyranose oxidase from the white rot fungus Irpex lacteus and its cooperation with laccase in lignin degradation. Process Biochem 49:2191–2198.
Baron R, Riley C, Chenprakhon P, thotsaporn K, Winter RT, Alfieri A, Forneris F, van Berkel WJH, Chaiyen P, Fraaije MW, Mattevi A, McCammon JA. 2009. Multiple pathways guide oxygen diffusion into flavoenzyme active sites. Proc Nat Acad Sci USA 106:10603-10608
Brown ME, Barros T, Chang MCY. 2012. Identification and Characterization of a Multifunctional Dye Peroxidase from a Lignin-Reactive Bacterium. ACS Chem Biol 7:2074–2081.
Brown ME, Chang MCY. 2014. Exploring bacterial lignin degradation. Curr Opin Chem Biol 19:1–7.
Cavener DR. 1992. GMC oxidoreductases: A newly defined family of homologous proteins with diverse catalytic activities. J Mol Biol 223:811–814.
Colpa DI, Fraaije MW, van Bloois E. 2014. DyP-type peroxidases: a promising and versatile class of enzymes. J Ind Microbiol Biotechnol 41:1–7.
Cragg SM, Beckham GT, Bruce NC, Bugg TDH, Distel DL, Dupree P, Etxabe AG, Goodell BS, Jellison J, McGeehan JE, McQueen-Mason SJ, Schnorr K, Walton PH, Watts JEM, Zimmer M. 2015. Lignocellulose degradation mechanisms across the Tree of Life. Curr Opin Chem Biol 29:108–119.
de Gonzalo G, Colpa DI, Habib MHM, Fraaije MW. 2016. Bacterial enzymes involved in lignin degradation. J Biotechnol 236:110–119.
Janusz G, Pawlik A, Sulej J, Świderska-Burek U, Jarosz-Wilkolazka A, Paszczyński A. 2017. Lignin degradation: Microorganisms, enzymes involved, genomes analysis and evolution. FEMS Microbiol Rev 41:941–962.
Kracher D, Scheiblbrandner S, Felice AKG, Breslmayr E, Preims M, Ludwicka K, Haltrich D, Eijsink VGH, Ludwig R. 2016. Extracellular electron transfer systems fuel cellulose oxidative degradation. Science 352:aaf3165.
Leitner C, Volc J, Haltrich D. 2001. Purification and Characterization of Pyranose Oxidase from the White Rot Fungus Trametes multicolor Purification and Characterization of Pyranose Oxidase from the White Rot Fungus Trametes multicolor 67:3636–3644.
Min K, Gong G, Woo HM, Kim Y, Um Y. 2015. A dye-decolorizing peroxidase from Bacillus subtilis exhibiting substrate-dependent optimum temperature for dyes and β-ether lignin dimer. Sci Rep 5:8245.
Pollegioni L, Tonin F, Rosini E. 2015. Lignin-degrading enzymes. FEBS J 282:1190–1213.
Qin X, Luo H, Zhang X, Yao B, Ma F, Su X. 2018. Dye-decolorizing peroxidases in Irpex lacteus combining the catalytic properties of heme peroxidases and laccase play important roles in ligninolytic system. Biotechnol Biofuels 11:302.
Singh R, Eltis LD. 2015. The multihued palette of dye-decolorizing peroxidases. Arch Biochem Biophys 574:56–65.
Sützl L, Laurent CVFP, Abrera AT, Schütz G, Ludwig R, Haltrich D. 2018. Multiplicity of enzymatic functions in the CAZy AA3 family. Appl Microbiol Biotechnol 102:2477–2492.
Wang F-F, Huang F, Ai M-Q. 2019 Synergetic Depolymerization of Aspen CEL by Pyranose 2-Oxidase and Lignin-degrading Peroxidases. BioResources 14:3481-3494
Wongnate T, Chaiyen P. 2013. The substrate oxidation mechanism of pyranose 2-oxidase and other related enzymes in the glucose-methanol-choline superfamily. FEBS J 280:3009–3027.
