Design of a minimal enzyme set for lignin depolymerization
JOINT-SUPERVISORS: Clemens PETERBAUER and Michael SAUER
Background.
Lignin makes up a significant fraction of woody biomass (25 – 35%) and is as such among the most abundant polymers on earth (Bugg et al., 2011; Varman et al., 2016). It is a complex aromatic heteropolymer with a variety of ether and C-C linkages, evolved to confer durability to plant structures and thus for a slow biodegradability. Only few microorganisms are able to degrade lignin into smaller units, most of them basidiomycete fungi, which grow slowly and are difficult to handle from an industrial point of view. Using lignocellulose (non edible plant material) as a resource, the lignin component is currently mostly burnt for energy, notwithstanding that utilization and valorization are seen as key factors to make biorefineries economically competitive (Varman et al., 2016; Lee at al. (2019); Shin et al. (2019). Hence, controlled depolymerization of lignin is a highly attractive challenge.
Research on lignin depolymerization has focused on fungal systems thus far (Pollegioni et al., 2015; Janusz et al., 2017). White-rot fungi produce heme-dependent lignin, manganese and versatile peroxidases as well as copper-dependent laccases and a peroxide-providing system of Auxiliary Activities Family 3 and 5 (AA3, AA5; Janusz et al., 2017), which is required for the peroxidase reaction cycle. AA3 enzymes were shown to work synergistically not only with peroxidases, but also with laccases (Ai et al., 2014; Wang et al., 2019). These enzyme systems are complicated to analyze and even more complicated to produce or use on industrial level. Consequently, no commercial biocatalytic process for lignin depolymerization is established to date (Bugg et al., 2011; Varman et al., 2016).
Recently, lignin oxidation was also observed in a number of soil bacteria, which seem to produce less complex enzyme systems attacking lignin (Cragg et al., 2015), with less redundancy. A number of activities have been described, among them small laccases from Streptomyces spp. (Majumdar et al., 2014), dye-decolorizing peroxidases (DyP, Brown et al., 2012; Brown and Chang, 2014) and pyranose oxidases, the sole AA3 enzymes identified in bacteria (Herzog and Sützl et al., 2019). It is largely unknown so far if these bacterial systems are able to degrade high molecular weight lignin in the absence of helping fungal enzymes, and which are the possible degradation products, or if they rely on a pre-degradation by other organisms.
Aims and methods.
This project aims at the characterization of selected bacterial enzymes as well as combinations of enzymes implicated in lignin degradation. The type of lignin molecules, which can be attacked, shall be characterized, as well as the products. The selected enzymes shall be investigated for their suitability for industrial use with the aim to obtain a high rate of depolymerization of lignin with a minimum of heterologously produced enzymes (“as few as possible, as many as required”).
The following enzymes shall be analyzed: The dye-decolorizing peroxidase DyP2 from Amycolatopsis sp. 75iv2 (Brown et al., 2012) as the only characterized representative from Actinomycetes thus far (Fig. 1). Corresponding genes are identified in the genomes of Streptomyces spp. and in related genera such as Kitasatospora, and a limited number will be included. We will further include BsDyP from B. subtilis (Min et al., 2015). Laccase: examples will be selected from Streptomyces coelicolor, Streptomyces lividans, Streptomyces viridosporus and Amycolatopsis sp. 75iv2, as described by Majumdar et al. (2014). Peroxide-providing enzymes: Kitasatospora aureofaciens POx (Herzog and Sützl et al., 2019), as well as perhaps similar enzymes characterized in a current project.
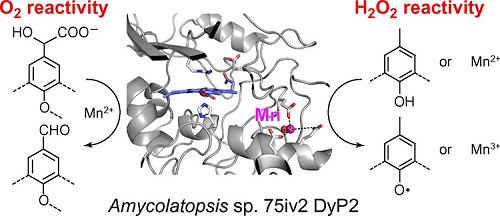
Fig. 1. Graphic representation of Mn2+-dependent oxidase activity and peroxidase activity of the Amycolatopsis sp. 75iv2 DyP2 (from Brown et al., 2012).
To allow an individual as well as rational combinatorial analysis of the enzymes, they will be heterologously expressed in Bacillus subtilis, as this is an established expression organism for a variety of industrial enzymes. It is amenable to genetic manipulation and a gram-positive microorganism, phylogenetically not too far from Streptomyces. (Harwood and Cranenburgh, 2008; van Dijl and Hecker, 2013; Cai et al., 2019). Laccases for lignin degradation have already been produced in B. subtilis (Shin, 2019) proving the suitability of this bacterium, and a surface display technology, which will be helpful in this context, has been developed (Anderson et al., 2011). A strain with a disrupted endogenous BsDyP-encoding gene will be used to avoid interference with the analysed enzymes. Finally, B. subtilis bears potential as cell factory for the valorization of lignin as it has been reported to metabolize certain aromatic lignin constituents (Lee et al., 2019; Shin et al., 2019; references therein).
Selected enzymes will be combined in “mini-operon” expression constructs containing a peroxidase and a pyranose oxidase, a laccase and a pyranose oxidase, as well as all three enzyme classes.
The obtained strains will be cultivated under defined conditions in bioreactors. This allows to produce and test the enzymes on a variety of lignin types and model substrates in various environmental conditions. The products will be analyzed by HPLC and selected LC-MS methods gaining deeper insight into the function of the enzymes.
Objectives
Functional expression of individual potentially ligninolytic enzymes in B. subtilis. (Peterbauer)
Cultivation of the strains under defined conditions in bioreactors in presence of lignin or lignin like model structures and detailed analysis of the supernatant thus analyzing the enzymatic activities in detail. (Sauer, collaboration with Hann)
Definition of a minimal set of enzymes for lignin degradation (Peterbauer, Sauer)
Construction of a B. subtilis based microbial cell factory for lignin degradation (Peterbauer)
Cultivation of the obtained cell factory on lignin or lignocellulose as sole carbon source and detailed analysis of the growth and production behaviour in bioreactors (Sauer)
Expected outcomes and impact
In how far soil bacteria are able to degrade lignin in absence of fungi is currently not known. Since cultivation of many of the strains in question is difficult, such analyses are complicated. This project will gain insight into the enzymatic capabilities of soil bacteria and lead to a better understanding of the natural carbon cycle. Furthermore, this knowledge will be useful for the construction of industrially useful cell factories for lignin valorization. While this goal has been long pursued, the complexity of the fungal lignin degrading enzymes did not allow any useful exploitation so far. Bacterial enzyme systems are decisively simpler, so there is a plausible potential that they might be of great use in industrial contexts. If their mode of activity is able to tackle the complexity of natural (undegraded) lignin is not known, but the objective of this project is to figure this out.
Fig. 2. Symbolic representation of the project´s components
Collaborations This thesis is envisioned as a joint project between PETERBAUER (protein engineering and characterization) and SAUER (cultivations, utilization of metabolites). Collaborations within BioToP are planned with HANN (analytics).
BOKU, Department of Biotechnology, Muthgasse 18, A-1190 Vienna, Austria
Email: michael.sauer@boku.ac.at.
BOKU, Department of Food Sciences and Technology, Muthgasse 11, A-1190 Vienna, Austria
Email: clemens.peterbauer@boku.ac.at.
REFERENCES:
Ai M-Q, Wang F-F, Zhang Y-Z, Huang F. 2014. Purification of pyranose oxidase from the white rot fungus Irpex lacteus and its cooperation with laccase in lignin degradation. Process Biochem 49:2191–2198
Anderson, T.D., Robson, S.A., Jiang, X.W., Malmirchegini, G.R., Fierobe, H.P., Lazazzera, B.A., Clubb, R.T., 2011. Assembly of minicellulosomes on the surface of Bacillus subtilis. Appl. Environ. Microbiol. 77 (14), 4849–4858.
Brown ME, Barros T, Chang MCY. 2012. Identification and Characterization of a Multifunctional Dye Peroxidase from a Lignin-Reactive Bacterium. ACS Chem Biol 7:2074–2081
Brown ME, Chang MCY. 2014. Exploring bacterial lignin degradation. Curr Opin Chem Biol 19:1–7
Bugg TDH, Ahmad M, Hardiman EM, Singh R (2011) The emerging role for bacteria in lignin degradation and bio-product formation. Curr Opin Biotechnol 22:394-400
Cai D, Rao Y, Zhan Y, Wang Q, Shen S (2019) Engineering Bacillus for efficient expression of heterologous protein: current progress, challenge and prospect. J Appl Microbiol 126:1632-1642
Cragg SM, Beckham GT, Bruce NC, Bugg TDH, Distel DL, Dupree P, Etxabe AG, Goodell BS, Jellison J, McGeehan JE, McQueen-Mason SJ, Schnorr K, Walton PH, Watts JEM, Zimmer M. 2015. Lignocellulose degradation mechanisms across the Tree of Life. Curr Opin Chem Biol 29:108–119
Harwood CR, Cranenburgh R (2008) Bacillus protein secretion: an unfolding story. Trends Microbiol 16:73-79
Herzog PL, Sützl L, Eisenhut B, Maresch D, Haltrich D, Obinger C, Peterbauer CK (2019) Versatile oxidase and dehydrogenase activity of bacterial pyranose 2-oxidase facilitates redox cycling with manganese peroxidase in vitro. Appl Env Microbiol 85:e00390-19
Janusz G, Pawlik A, Sulej J, Świderska-Burek U, Jarosz-Wilkolazka A, Paszczyński A. 2017. Lignin degradation: Microorganisms, enzymes involved, genomes analysis and evolution. FEMS Microbiol Rev 41:941–962
Lee S, Kang M, Bae J-H, Sohn J-H, Sung BH (2019) Bacterial Valorization of Lignin: Strains, enzymes, Conversion Pathways, Biosensors and Perspectives. Frontiers Bioeng Biotechnol 7:209
Majumdar S, Lukk T, Solbiati JO, Bauer S, Nair SK, Cronan JE, Gerlt JA (2014) Roles of Small Laccases from Streptomyces in Lignin Degradation. Biochemistry 53: 4047-4058
Min K, Gong G, Woo HM, Kim Y, Um Y. 2015. A dye-decolorizing peroxidase from Bacillus subtilis exhibiting substrate-dependent optimum temperature for dyes and β-ether lignin dimer. Sci Rep 5:8245.
Pollegioni L, Tonin F, Rosini E. 2015. Lignin-degrading enzymes. FEBS J 282:1190–1213
Shin SK, Ko YJ, H JE, Han SO (2019) Studies of advanced lignin valorization based on various types of lignolytic enzymes and microbes. Biores Technol 289:121728
Van Dijl JM, Hecker M (2013) Bacillus subtilis: from soil bacterium to super-secreting cell factory. Microb Cell Fact 12:3
Varman AM, He L, Follenfant R, Wu W, Wemmer S, Wrobel SA, Tang YJ, Singh S (2016) Decoding how a soil bacterium extracts building blocks and metabolic energy from ligninolysis provides road map for lignin valorization. Proc Nat Acad Sci USA 113: E5802-E5811
Wang F-F, Huang F, Ai M-Q. 2019 Synergetic Depolymerization of Aspen CEL by Pyranose 2-Oxidase and Lignin-degrading Peroxidases. BioResources 14:3481-3494
