Understand and exploit sigma-appropriation as a tool for transcription control in E. coli
SUPERVISOR: Reingard GRABHERR
Background.
Control of transcription is crucial for correct gene expression and orderly development of a healthy cell or organism. During an infectious cycle of viruses, the host’s synthesis machinery is taken over and transcription and translation is redirected towards synthesis of viral components. There exist many different strategies used by viruses and phages to force the cellular transcriptional machinery into preferring viral promoters and to downregulate or stop the host’s transcription. The T7 phage brings its own RNA polymerase that recognizes T7 phage promoters only and thereby provides T7 specific mRNA production. At the same time phage proteins are being expressed that interfere with the host’s RNA polymerase, which leads to a shut-down of E. coli gene expression. In contrast to this strategy, the T4 phage captures and reshapes the E. coli RNA polymerase thereby redirecting it to T4 phage promoters (Hinton, 2010), a process which is called sigma appropriation. The mechanism is described as interaction of two phage encoded proteins, AsiA and MotA with the E. coli sigma-factor 70 (James et al., 2016). This interaction changes the structure of the sigma-factor so that binding to the -35 region of host promoters is prevented and binding to T4 phage middle promoters is enhanced. The N-terminal domain of MotA is responsible to bind to the MotA-box, a motif that is only present in T4 phage middle promoters. Interaction studies by James et al. (2016) and Shi et al. (2019) investigated the crucial domains for binding to the respective promoters.
Transcription control in E. coli is also a required tool in biotechnology for recombinant protein expression. Many expression vectors are based on the T7 system, where the combined presence of the T7 promoter and T7 polymerase within the expression host leads to high level expression. Many other inducible systems exist as well as antisense RNA based strategies for down-regulation of transcription or targeted gene deletion for complete knock-out (lee et. Al). Phage strategies for interference with the host RNA polymerase have also successfully been incorporated into innovative and efficient E. coli-based expression systems (Mairhofer et al.). Yet, a molecular tool for targeted up-regulation of a certain cellular gene, without having to genetically engineer the respective promoter is unavailable. For improved recombinant protein expression in E. coli (e.g. by higher co-expression of chosen chaperones), as well as for all systems biology approaches where genes need to be upregulated in a high through-put format, methods for increasing the interaction of the cellular RNA-polymerase and a chosen promoter would be very helpful.
Although, the structural basis of sigma-appropriation has been studied in detail (Shi et al., 2019) not all functions of the participating components have been fully understood. AsiA is a protein consisting of 90 amino acids. Expression of AsiA wildtype is toxic in E. coli, however there exists a set of deletion and substitution mutants that allow for expression and are still functional for initiating transcription, whenever MotA is present (Pal et al., 2003). Starting with these mutants, we would first test the effect on our E. coli strains HMS174 and BL21 in the presence and absence of MotA. MotA is responsible for binding to the MotA-box which is present in T4 phage promoters. In our strains, there are no T4 phage promoters or MotA motifs and thus, we need an alternative strategy to guide AsiA to a chosen promoter. We will test interaction of the AsiA sigma 70 complex, either with or without co-expression of MotA, with guide RNAs as it is used in CRISPR-Cas9 based methods. Here a guide RNA that is homologous to a specific DNA sequence, is recognized and bound by the endonuclease Cas9 and thereby brought into close distance to the specific genome region. The same strategy will be used to guide AsiA to a chosen promoter, whereby transcription initiation should be enhanced.
Aims and methods.
There exists a variety of molecular biology tools for targeted transcription control in E. coli, e.g. inducible promoters and phage T7 based expression systems. Following the strategy used by the T4 phage, our goal is to capture the E. coli RNA polymerase and bring it to chosen cellular promoters by an artificial guide RNA in order to up-regulate chosen genes without genetic manipulation thereof. We plan to modify the T4 phage protein AsiA in a way, that it is no longer toxic to E. coli. Various mutants of AsiA (Pal et al. 2003) will be expressed in E. coli strains HMS174 and BL21 with and without co-expression of MotA. E. coli strains will be characterized in terms of growth and recombinant protein expression of a model product. We further will construct fusion proteins between AsiA and the guide-RNA binding domain of Cas9 and repeat the experiments. Appropriate strains will be generated in order to test the possibility to specifically activate certain promoters by AsiA complexes via guide RNAs. The challenge is to maintain the structural requirements of AsiA to interact with sigma factor 70, therefore, protein structure modeling in collaboration with Chris Oostenbrink will be included. Based on these results it might be necessary to design and test additional mutants. In order to test various AsiA mutants and combinations with MotA we will construct monitoring strains containing the reporter gene GFP (Figure 1). Four different E. coli strains for HMS174 as well as for BL21 will be generated by integrating a GFP expression cassette into the chromosome. Four different promoters will be tested, (i) a T4 middle promoter, which contains the MotA-box and should be recognized whenever MotA is present in the final protein complex, (ii) a weak inducible promoter (e.g. arabinose inducible promoter), that can be turned on by addition of an inducer, (iii) an E. coli promoter (e.g. of a chaperone) and (iv) an artificial sequence that is free of any promoter specific motifs. Two different types of plasmids will be constructed expressing non-toxic AsiA mutants fused to the RNA binding domain of Cas9 and encoding for a guide RNA specific for the respective promoter region. Based on this experimental set-up various versions and structures of AsiA can be tested in terms of promoter activation, in order to elucidate some of the functional mechanisms as well as providing a novel systems biology tool.
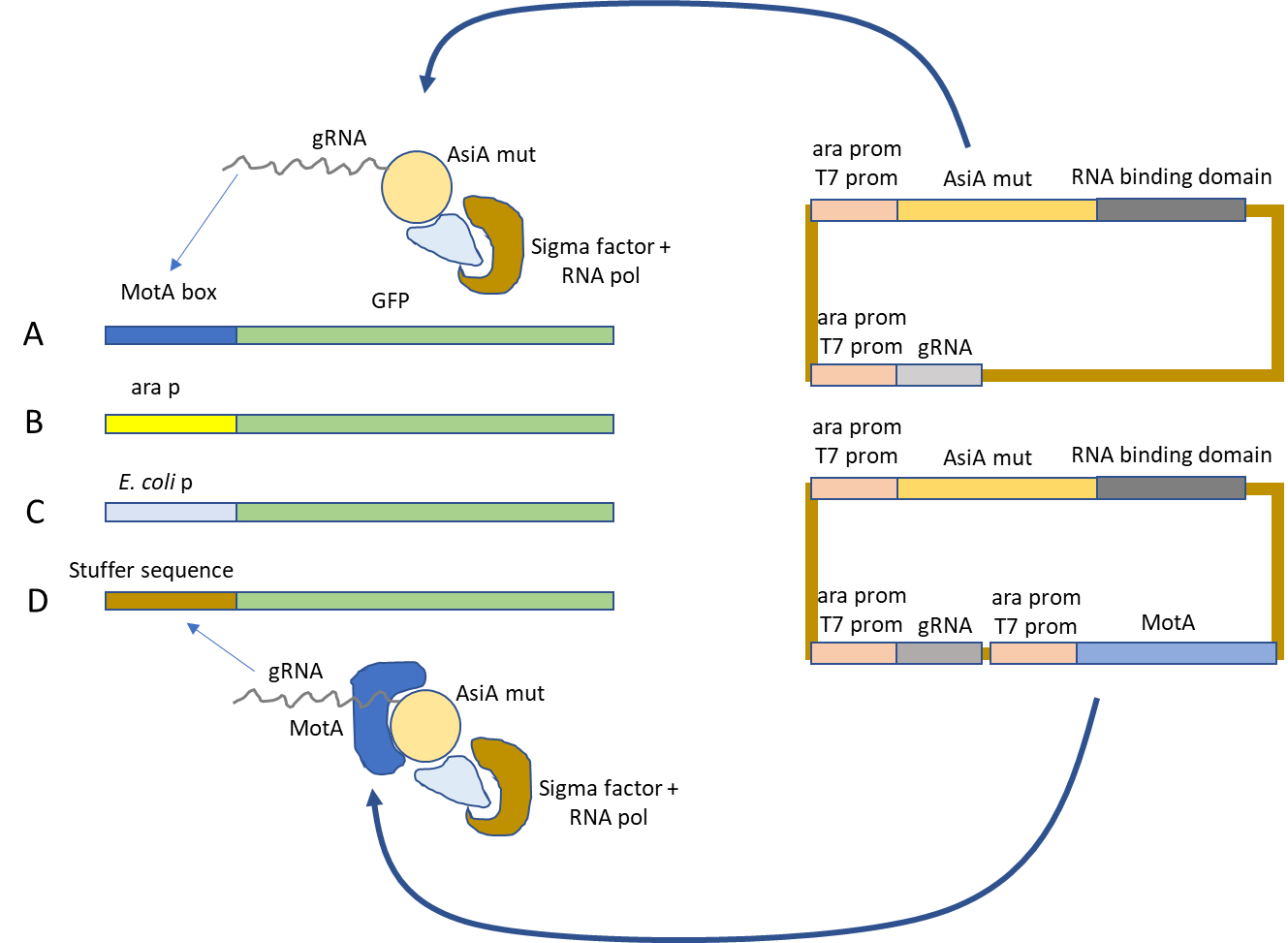
Fig. 1.Four different E. coli strains will be generated by insertion of a GFP expression cassette into the host’s genome: A: promoter containing a MotA box, B: containing the arabinose inducible promoter ara p. C: containing an E. coli promoter, D: containing a stuffer sequence without promoter specific sequences. Two different plasmids will be used to transform each of the four strains, one expressing AsiA fused to the RNA binding domain of Cas9 under control of the weak ara p or the strong T7 promoter and the promoter specific guide RNA under control of the weak ara p or the strong T7 promoter and a second plasmid containing the same expression cassettes plus MotA under control of the ara p or the T7 promoter.
In our experimental set-up, E. coli production strains contain expression cassettes encoding Fab fragments integrated in their genome. These strains will be equipped with the previously optimized sigma appropriation complex (consisting of AsiA mutant fused to RNA binding protein, MotA if required, under control of a feasible promoter and integrated in the E. coli genome). Various guide RNAs targeting a set of chosen genes will be co-expressed during a fermentation process. The overall goal is to enhance yield and quality of the Fab fragment by up-regulation of one or more co-factors and to provide a stable and robust E. coli-based production process.
James D. T., Cardozo, T., Abell, L. E., Hrieh, M.-L., Miller Jenkins, L. M., Jha, S. S., Hinton, D. M. (2016) Visualizing the phage T4 activated transcription complex of DNA and E. coli RNA polymerase. Nucleic Acids research 44, doi: 10.1093/nar/gkw656
Kutter, E., Bryan, D., Ray, G., Brewster, E., Blasdel, B., Burton, G. (2018) From Host to Phage metabolism: Hot Tales of Phage T4’s Takeover of E. coli.
Hinton, D. M. (2010) Transcriptional control in the prereplicative phase of T4 development. Virology Journal 7
Shi, J., Wen, A., Zhao, M., You, L., Zhang, Y., Feng, Y. (2919) Structural basis of sigma appropriation. Nucleic Acids Research 47, 9423-9432.
