Engineering the interface of Fab fragments for the generation of bispecific antibodies
SUPERVISOR: JOHANNES GRILLARI
Background.
Targeting two different types of antigens with a single molecule has rendered bispecific antibodies an attractive concept in cancer therapy. In contrast to normal monospecific antibodies bispecific IgGs exhibit two different specificities, one on each Fab arm. This feature offers the possibility to link a cancer antigen displayed on a tumour cell with an antigen displayed by a cytotoxic T cell (e.g. CD3). Engineering a bispecific antibody has proven to be a difficult task. A bispecific IgG consists of two different heavy and two different light chains, however, up to a certain degree, the pairing of the chains occurs randomly. Several technologies have been developed to overcome the homodimerisation of the heavy chains, e.g. knobs-into-holes (KiH, Ridgway et al. 1996) and strand-exchange engineered domain (SEED, Davis et al. 2010). Nevertheless, the pairing of the light to the heavy chains still occurs randomly which makes production inefficient and purification of the antibody cumbersome. At the Christian Doppler Laboratory for antibody engineering we aim to engineer the interface within the constant region of the Fab arm, consisting of the domains CL and CH1.
Aims and methods.
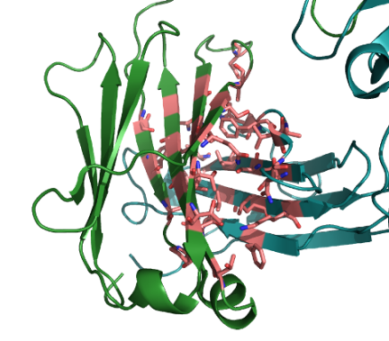
IIn order to allow specific pairing between heavy and light chains we aim to engineer the interface between the domains CH1 and Ck on one Fab arm (see picture, Ck in green, CH1 in blue, possible interface residues highlighted in red) to allow only the pairing with the accordingly engineered opposite chain. At the same time, the undesired pairing to the either wildtype or differently engineered opposite chain in the other Fab arm shall be reduced to a minimum. Several mutations which seem to weaken the binding to the opposite wildtype chain have been identified so far using either a yeast display system with covalent linkage of the Fab to the yeast cell surface or the expression of full IgGs in HEK 293 cells. Mutations that restore the interface will be identified either by a rational mutagenesis or a library approach using a novel display system on HEK 293 cells. The resulting bispecific antibodies will be produced and their expression level, thermostability, folding status, antigen binding affinity and purity will be determined and further optimised.
Davis, J. H., Aperlo, C., Li, Y., Kurosawa, E., Lan, Y., Lo, K.-M., Huston, J. S. (2010) SEEDbodies: fusion proteins based on strand-exchange engineered domain (SEED) CH3 heterodimers in an Fc analogue platform for asymmetric binders or immunofusions and bispecific antibodies. Protein engineering, design and selection: PEDS, 23(4)
Ridgway, J. B., Presta, L. G., Carter, P. (1996) Knobs-into-holes engineering of antibody CH3 domains for heavy chain heterodimerization. Protein engineering, 9(7), 617–21.
