Interrelation of core metabolism with the production of heterologous proteins in yeast
SUPERVISOR: MATTANOVICH DIETHARD
Project assigned to: JUSTYNA NOCON
Background.
It has been observed in numerous cases that the expression of heterologous proteins in microorganisms decreases growth and biomass yield, even when the absolute amounts of the produced protein is quantitatively negligible in relation to biomass and total cellular protein synthesized (Mattanovich et al., 2004). Especially when producing secretory proteins in yeasts, the total amounts of product are usually very low compared to the produced total cellular protein, but still leads to a significant burden on the host cells (Goergens et al., 2001; Vigentini et al., 2005; Ramon et al., 2007; Baumann et al., 2008; Dragosits et al., 2009). The reasons behind this effect are largely unknown. Up to now it is only speculation whether heterologous protein production, and particularly secretion, lead to futile metabolic cycles, or other substrate consuming metabolic processes so that finally the biomass yield decreases significantly.
In order to understand the metabolic processes behind this phenomenon on a quantitative level, the flux rates of substrate (carbon and energy source) to biomass, energy production and redox equivalents need to be elucidated. The method of choice is metabolic flux analysis employing isotope labelling, in combination with quantitative analysis of specific metabolites and co-factors. Metabolic flux analysis has been established for P. pastoris by the group of Pau Ferrer (Sola et al., 2004; Sola et al., 2007). In collaboration with this group we are currently setting up metabolic flux analysis employing LC-MS for isotope analysis (Wiechert, 2001) and flux balance calculation based on open-source software like OpenFLUX (Queck et al., 2009). Metabolic flux analysis will allow for an understanding of the divergent pathways of carbon flux from the substrate to biomass, (by)products, and CO2, providing insight at which steps of the cellular metabolism an additional drain exists leading away from biomass synthesis.
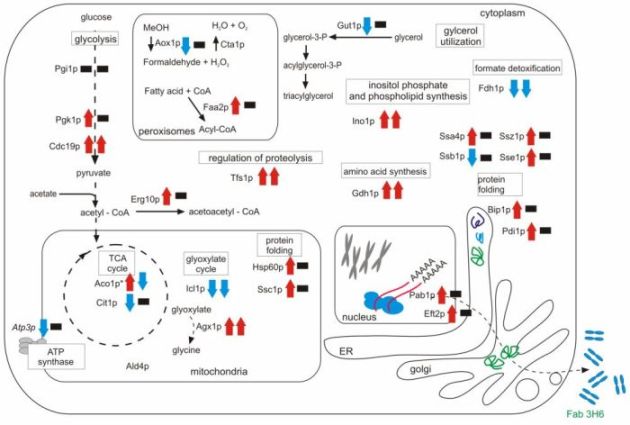
During recent projects we have accumulated strong evidence that redox processes – especially concerned with disulfide bond formation in heterologous proteins – may play a major role in the metabolic burden exerted by heterologous protein synthesis in yeast (Gasser et al., 2006; Gasser et al., 2007; Gasser et al., 2008). Therefore it appears imperative to investigate the redox system of the cellular systems as well. Preliminary experiments of the intracellular proteome of P. pastoris indicate that several enzymes involved in the TCA cycle, amino acid biosynthesis and pentose phosphate shunt show different abundance between a recombinant protein producing strain and its wild-type control (Dragosits et al., 2009). Recently, similar results have been obtained for S. cerevisiae expressing an intracellular recombinant protein, where genes of the central metabolic pathways are downregulated in the production strain (Diaz et al., 2009).
Aims and methods.
Based on already established production strains for complex heterologous proteins (antibody fragments – Gasser et al., 2006; trypsinogen – Hohenblum et al., 2004) and their respective non-producing control strains, the carbon fluxes to precursors, energy production and redox equivalents shall be quantified. In addition, the analysis of redox equivalents (NAD+/NADH, NADP+/NADPH) and the major redox buffer system (oxidized and reduced glutathione) will allow for understanding of the correlation between the central flux and redox maintenance.
Metabolic flux analysis will be based on 13C labelling in chemostat cultures, comparing expression strains with their isogenic non-producing controls. Following appropriate quenching and extraction, isomers will be quantified by LC-MS/MS. Reduced and oxidized glutathione will be measured by LC-TOFMS, and NAD/NADP nucleotides by LC-MS/MS. Measuring by-product formation, off-gas and total biomass will allow for a total carbon balance. Quantification of the heterologous protein produced is important in order to correlate its production to the respective impact on central metabolism.
The final aim of the project is to build a model of the specific processes linking the core metabolism to heterologous protein production based on the metabolic rates of carbon flux, energy and redox equivalents.
Baumann, K., Maurer, M., Dragosits, M., Cos, O., Ferrer, P., Mattanovich, D. (2008) Hypoxic fed-batch cultivation of Pichia pastoris increases specific and volumetric productivity of recombinant proteins. Biotechnol. Bioeng. 100, 177-183
Díaz, H., Andrews, B. A., Hayes, A., Castrillo, J., Oliver, S., Asenjo, J. A. (2009) Global gene expression in recombinant and non-recombinant yeast Saccharomyces cerevisiae in three different metabolic states. Biotechnol. Adv., in press
Dragosits, M., Stadlmann, J., Albiol, J., Baumann, K., Maurer, M., Gasser, B., Sauer, M., Altmann, F., Ferrer, P., Mattanovich, D. (2009). The effect of temperature on the proteome of recombinant Pichia pastoris. J. Proteome Res. 8, 1380-1392
Gasser, B., Maurer, M., Gach, J., Kunert, R., Mattanovich, D. (2006) Engineering of Pichia pastoris for improved production of antibody fragments. Biotechnol. Bioeng. 94, 353-361
Gasser, B., Sauer, M., Maurer, M., Stadlmayr, G., Mattanovich, D. (2007) Transcriptomics based identification of novel factors enhancing heterologous protein secretion in yeasts. Appl. Environ. Microbiol. 73, 6499-6507
Gasser, B., Saloheimo, M., Rinas, U., Dragosits. M., Rodríguez-Carmona, E., Baumann, K., Giuliani, M., Parrilli, E., Branduardi, P., Lang, C., Porro, D., Ferrer, P., Tutino, M. L., Mattanovich, D., Villaverde, A. (2008) Protein folding and conformational stress in microbial cells producing recombinant proteins: a host comparative overview. Microb. Cell Fact. 7, 11
Görgens, J. F., van Zyl, W. H., Knoetze, J. H., Hahn-Hägerdal. B. (2001) The metabolic burden of the PGK1 and ADH2 promoter systems for heterologous xylanase production by Saccharomyces cerevisiae in defined medium. Biotechnol. Bioeng. 73, 238-245
Hohenblum, H., Gasser, B., Maurer, M., Borth, N., Mattanovich, D. (2004) Effects of gene dosage, promoters and substrates on unfolded protein stress of recombinant Pichia pastoris. Biotechnol. Bioeng. 85, 367-375
Mattanovich, D., Gasser, B., Hohenblum, H., Sauer, M. (2004) Stress in recombinant protein producing yeasts. J. Biotechnol. 113, 121-135
Ramón, R., Ferrer, P., Valero, F. (2007) Sorbitol co-feeding reduces metabolic burden caused by the overexpression of a Rhizopus oryzae lipase in Pichia pastoris. J. Biotechnol. 130, 39-46
Solà, A., Maaheimo, H., Ylönen, K., Ferrer, P., Szyperski, T. (2004) Amino acid biosynthesis and metabolic flux profiling of Pichia pastoris. Eur. J. Biochem. 271, 2462-2470
Solà, A., Jouhten, P., Maaheimo, H., Sánchez-Ferrando, F., Szyperski, T., Ferrer, P. (2007) Metabolic flux profiling of Pichia pastoris grown on glycerol/methanol mixtures in chemostat cultures at low and high dilution rates. Microbiology 153, 281-290
Vigentini, I., Brambilla, L., Branduardi, P., Merico, A., Porro, D., Compagno, C. (2005) Heterologous protein production in Zygosaccharomyces bailii: physiological effects and fermentative strategies. FEMS Yeast Res. 5, 647-652
Wiechert, W. (2001) 13C metabolic flux analysis. Metab Eng. 3, 195-206
Quek, L. E., Wittmann, C., Nielsen, L. K., Krömer, J. O. (2009) OpenFLUX: efficient modelling software for 13C-based metabolic flux analysis. Microbial Cell Fact. 8, 25
