Assembly and interaction mechanisms of S-layer glycoproteins for applications in nanobiotechnology
SUPERVISOR: PAUL MESSNER
Project assigned to: ZOE MEGSON
Background.
S-Layers represent the outermost cell envelope component of many Archaea and Bacteria. Previous microscopical analyses have shown that S-layers form crystalline lattices which cover the cells completely, revealing first insights into the self-assembly process (Sleytr et al., 2007). Self-assembly of molecules into crystalline arrays is a new and rapidly growing transdisciplinary scientific and engineering field. S-layers are periodic structures which exhibit identical physicochemical properties on each molecular unit down to the subnanometer scale. Functional groups are aligned on the S-layer lattice in well-defined positions and orientations. Chemical analyses of different S layer proteins have demonstrated that glycosylation is the most frequent modification of S layer proteins. Generally, cell surface glycoconjugates play critical roles in interactions between prokaryotes and their immediate environment(s). It is particularly the glycan portion which is involved in biological processes, including involvement in many cellular processes, such as recognition, signalling, trafficking, biological half-life time, and adhesion events. Due to the general benefits of a protein self-assembly system that is able to form regularly-arrayed two-dimensional matrices with nanometer precision, S-layer glycoprotein-carrying organisms, where the carbohydrate portion protrudes off the protein crystal, are promising candidates for applications in the fields of nanobiotechnology and biomedicine. By genetic engineering the production of rationally-designed, biofunctional S-layer neoglycoproteins (Steiner et al., 2008) and glycopeptides, as well as in vivo cell surface glycan display is feasible.
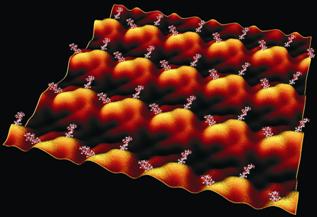
For this endeavour, the Gram-negative bacterium Tannerella forsythia and the Gram-positive organism Paenibacillus alvei CCM 2051T, both of which are covered by S layer glycoproteins, are serving as models. The importance of T. forsythia lies in the documented virulence potential of its S layer. While S layer glycosylation of this organism remains to be elucidated, the S layer glycan of the second model organism is already known. The benefit of this organism is its genetic transformability (Zarschler et al., 2009), while different other S-layer glycoprotein-carrying Gram-positive organisms have resisted to uptake foreign DNA. This makes it a prime candidate for in vivo glycan display approaches. For either organism, the assembly properties of the S-layer glycoprotein are only poorly understood and no crystallization experiments have been performed in the past.
Aims and methods.
Since the S-layer properties of both organisms are only superficially characterized, the principal understanding of the assembly process of the S-layer (glyco)proteins are a prerequisite for any nanobiotechnological application (Schäffer et al., 2007). New analysis methods such as scanning probe microscopy (SPM) (Hinterdorfer et al., 1996), surface plasmon resonance (SPR) spectroscopy, quartz crystal microbalance measurements with dissipation monitoring (QCM-D) (Cooper and Singleton, 2007), or high resolution magic angle spinning NMR (HR MAS-NMR) measurements (Händel et al., 2003) (of labeled proteins) will allow improvement of the general understanding of the assembly process of S-layer (glyco)proteins and their interactions with ligands, including the identification of possible receptors involved in S-layer binding to cells.
S-layer proteins are able to form self-assembly products in suspension and to reassemble at interfaces. In particular, the formation of monolayers on solid supports (e.g. silicon surfaces, mica or gold), allows studying structural and functional parameters by SPM. Based on well established methods for imaging biomolecules (proteins, lipids, etc.) by high resolution SPM, single-molecule force spectroscopy in vitro will enable the characterization of the crystallization properties at the nanometer scale. Although insights into protein-binding pathways or binding potentials of receptor-ligand pairs have already been obtained experimentally, the molecular mechanisms underlying the mechanical functions can only be studied through molecular dynamics simulations with atomic structural models (“in silico”). Force spectroscopy experiments and steered molecular dynamic simulations in which external forces are used to study response and function of the mentioned S layer proteins will be performed. The basic idea is to establish the reassembly of the S-layer proteins on flat solid supports, record force-distance curves between tip and S-layers and investigate the force-induced conformational changes by computer analysis. The obtained results shall provide more insight into the molecular details of the reassembly process.
The interaction of the S-layer model systems with corresponding cell wall components, but also with other components like, for instance, proteins, pathogens, and cells will be investigated by two powerful surface-sensitive techniques: QCM-D and SPR. Both methods are ultra-sensitive mass sensors that monitor the real-time change in adsorbed amount of material. In contrast to SPR, QCM-D senses in addition to the adsorbed amount of material also the mass of trapped/coupled water on various surfaces and the viscoelastic properties of the film during adsorption. By presenting different surfaces to biomolecules, SPR and QCM-D can also differentiate between varying affinities of adsorption and kinetics in response to specific carbohydrate engineering techniques. Using both techniques in parallel, but also combined with SPM and HR MAS-NMR, is a powerful approach enabling interesting insights for the understanding of complex self-assembly processes, protein-carbohydrate interaction, the hydration of biomaterials, cell-cell interaction, biofilm formation, and anti-carbohydrate-antibody interaction.
Cooper, M. A., Singleton, V. T. (2007) A survey of the 2001 to 2005 quartz crystal microbalance biosensor literature: applications of acoustic physics to the analysis of biomolecular interactions. J. Mol. Recognit. 20, 154-184
Händel, H., Gesele, E., Gottschall, K., Albert, K. (2003) Application of HRMAS 1H NMR spectroscopy to investigate interactions between ligands and synthetic receptors. Angew. Chem. Int. Ed. 42, 438-442
Hinterdorfer, P., Baumgartner, W., Gruber, H. J., Schilcher, K., Schindler, H. (1996) Detection and localization of individual antibody-antigen recognition events by atomic force microscopy. Proc. Natl. Acad. Sci. USA 93, 3477-3481
Schäffer, C., Novotny, R., Küpcü, S., Zayni, S., Scheberl, A., Friedmann, J., Sleytr, U. B., Messner, P. (2007) Novel types of biocatalysts based on the S-layer self-assembly system from Geobacillus stearothermophilus NRS 2004/3a, exemplified with the glucose-1-phosphate thymidylyltransferase RmlA: a nanobiotechnological approach. Small 3, 1549-1559
Sleytr, U. B., Huber, C., Pum, D., Schuster, B., Ilk, N., Egelseer, E. M. (2007) Nanobiotechnology with S-layer proteins. FEMS Microbiol. Lett. 267, 131-144
Steiner, K., Hanreich, A., Kainz, B., Hitchen, P. G., Dell, A., Messner, P., Schäffer, C. (2008). Recombinant glycans on an S-layer self-assembly protein: a new dimension for nanopatterned biomaterials. Small 10, 1728-1740
Zarschler, K., Janesch, B., Zayni, S., Schäffer, C., Messner, P. (2009). Construction of a gene knockout system for application in Paenibacillus alvei CCM 2051T, exemplified by the S-layer glycan biosynthesis initiation enzyme WsfP. Appl. Environ. Microbiol. 75, 3077-3085
