Advanced molecular modeling tools to quantify protein-protein interactions
SUPERVISOR: CHRIS OOSTENBRINK
Project assigned to: MANUELA MAURER
Background.
The starting point of investigations for this project is the role of water molecules in biomolecular interactions. The periplasmic oligopeptide binding protein A (OppA), for example, is well-known for water-mediated ligand binding. OppA relies on a layer of ordered, interfacial water molecules in the active site (surrounding e.g. the side chains of bound peptides) to accommodate a broad range of ligands with different physico-chemical properties. Different water configurations have been observed for different oligopeptides, binding to the active site. This promiscuous binding of tripeptides to the OppA protein, combined with the highly flexible peptidic character of the ligands and the wealth of published experimental data, makes the OppA system an excellent test case for studying the relationships between structure and thermodynamics, as well as the (mediating) effect of solvent molecules, and furthermore for advanced computational approaches.
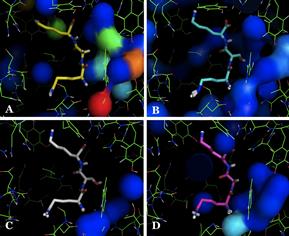
Aims and methods.
In this project we plan to focus on the use of enhanced sampling techniques, specifically the use of soft atoms and replica exchange methods. Advanced simulation schemes and judiciously chosen modifications to Hamiltonians have the potential to make free energy, enthalpy and entropy calculations tremendously more efficient, also for currently challenging biomolecular systems that involve large conformational changes.
Tools will need to be developed to determine which energetic barriers are responsible for slowly evolving conformational changes. Subsequently, these barriers can be specifically removed by modifying the Hamiltonian. The combination of intentionally non-physical potential energy surfaces (e.g. through soft atoms) and modern REMD simulations will prove to be highly useful for a wide range of applications. Using the OPPA system as an example application, quantitative, rigorous methods which include the individual contributions of the solvent and the loss of conformational flexibility of the protein and the ligands theoretically allows us to properly incorporate the effects of active site water molecules on the binding thermodynamics.
Initially, we aim to perform free-energy calculations for tripeptides binding to OppA, using replica exchange thermodynamic integration and one-step perturbation approaches. The small tripeptides allow us to develop generalized models for free energy calculations between amino acids, which can subsequently be used to compute protein-protein interactions. Additionally, the complex and changing water network in OppA requires extensive sampling, and it is therefore likely to profit significantly from enhanced sampling techniques. Through the use of these techniques we subsequently hope to achieve a more accurate description and understanding of functionally relevant water molecules in biomolecular interactions at large.
Li, Z.; Lazaridis, T.: Water at biomolecular binding interfaces. Phys. Chem. Chem. Phys. 2007, 9, 573-581.
Sleigh, S. H.; Seavers, P. R.; Wilkinson, A. J.; Ladbury, J. E.; Tame, J. R. H.: Crystallographic and calorimetric analysis of peptide binding to OppA protein. J. Mol. Biol. 1999, 291, 393-415.
Wang, T.; Wade, R. C.: Comparative binding energy (COMBINE) analysis of OppA-peptide complexes to relate structure to binding thermodynamics. J. Med. Chem. 2002, 45, 4828-4837.
Riniker, S.; Christ, C. D.; Hansen, H. S.; Huenenberger, P. H.; Oostenbrink, C.; Steiner, D.; van Gunsteren, W. F.: Calculation of relative free energies for ligand-protein binding, solvation, and conformational transitions using the GROMOS software. J. Phys. Chem. B 2011, 115, 13570-13577.
J.A. Garate and C. Oostenbrink: Free energy differences between states with different conformational ensembles. J. Comput. Chem. 2013, 34, 1398 – 1408.
J. Hritz and C. Oostenbrink: Efficient free energy calculations for compounds with multiple stable conformations separated by high energy barriers. J. Phys. Chem. B. 2009, 113, 12711-12720.
G. Nagy and C. Oostenbrink: Rationalization of Sterospecific Binding of Propanolol to Cytochrome P450 2D6 by Free Energy Calculations. Eur. Biophys. J. 2012, 41, 1065 – 1076.
