Glycoprofiling of oral pathogens – identification of novel glycoproteins and impact of glycosylation on biofilm formation
SUPERVISOR: CHRISTINA SCHÄFFER
Project assigned to: CORNELIA RATH
Background.
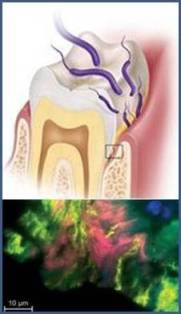 Multispecies biofilms develop in the human oral cavity on mucosal and dental surfaces with a precise spatial organization of interacting bacteria. Significant habitat changes due to food ingestion cause increased flow of gingival crevicular fluid and pH changes. Insufficient care can lead to a switch of the resident bacterial composition to putative pathogens responsible for periodontal diseases (Marsh et al., 2011)
Multispecies biofilms develop in the human oral cavity on mucosal and dental surfaces with a precise spatial organization of interacting bacteria. Significant habitat changes due to food ingestion cause increased flow of gingival crevicular fluid and pH changes. Insufficient care can lead to a switch of the resident bacterial composition to putative pathogens responsible for periodontal diseases (Marsh et al., 2011)
The oral Gram-negative anaerobic pathogen Tannerella forsythia (Tf) belongs, together with Porphyromonas gingivalis and Treponema denticola, to the so called “red-complex” of bacteria. Next to others, these bacteria form sub-gingival plaque biofilms leading to periodontitis. Tf is covered with a unique 2D crystalline glycosylated S(surface)-layer, built by two unique S-layer glycoproteins (TfsA-GP, TfsB-GP) (Sekot et al., 2012; Posch et al., 2011), which have been shown to clearly impact the bacterium’s ability to form biofilm (Honma et al., 2007). Increased biofilm formation has been observed for an isogenic Tf wecC mutant, due to the presence of truncated S-layer glycans (Posch et al., 2011). Furthermore, as unpublished data by our group demonstrate, biofilm architecture significantly changes in Tf mutants lacking one or both S-layer glycoproteins, with mutant cells forming biofilms with less aggregates, but with higher substrate affinity. This indicates that glycoproteins on the cell surface might serve as lead components for combating biofilm development, and, consequently, establishment of periodontal diseases (Posch et al., 2012).
Aims and methods.
We hypothesize that also other bacteria commonly present in dental plaque possess “S-layer-like” structures or glycoproteins attached to the cell surface, affecting the bacterial intra- and inter-species communication in the oral cavity and the capacity to interact with host cells. We carefully selected six candidates from the oral microflora, i.e., Capnocytophaga ochracea, Eubacterium nodatum, Fusobacterium nucleatum, Selenomonas sputigena, Streptococcus gordonii and Veillonella parvula,for glycobiology and biofilm studies. Identity and cellular location of selected candidate proteins will be investigated, as well as nature and structure of the attached glycans in order to extend the current knowledge of the glycobiology of oral pathogens. Defined glycosylation mutants will allow the assessment of the role of glycosylation in biofilm formation, and consequently, the establishment of virulence mechanisms of oral pathogens.
Marsh, P.D. et al. (2011) Dental plaque biofilms: communities, conflict and control. Periodont 2000, 55, 16-5.
Sekot, G. et al. (2012) Analysis of the cell surface layer ultrastructure of the oral pathogen Tannerella forsythia. Arch Microbiol 194, 525-539.
Posch, G. et al. (2011) Characterization and scope of S-layer protein O-glycosylation in Tannerella forsythia. J Biol Chem 286, 38714-38724.
Honma, K. et al. (2007) Role of a Tannerella forsythia exopolysaccharide synthesis operon in biofilm development. Microb Pathog 42, 156-166.
Posch, G. et al. (2012) Glycobiology aspects of the periodontal pathogen Tannerella forsythia. Biomolecules 2, 467-482.
