Human peroxidasins – new multidomain oxidoreductases in the vascular system
SUPERVISOR: CHRISTIAN OBINGER
Project assigned to: MONIKA SOUDI
Background.
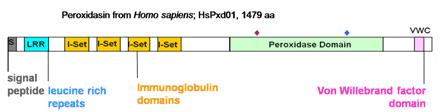
Recently, based on the peroxidase domain of mammalian peroxidases, we defined the peroxidase-cyclooxygenase superfamily and its subdivision in seven subfamilies (Zamocky et al., 2008). The second subfamily is represented by peroxidasins found in invertebrates and vertebrates including mammals. The first peroxidasin was found in Drosophila and described as a protein combining peroxidase and extracellular matrix motifs (Nelson et al., 1994). The multidomain proteins contain a signal peptide, several leucine-rich regions, followed by immunoglobulin domains, a linking region, the peroxidase domain, and a carboxy terminal van Willebrand type C protein-protein interaction. The peroxidase domains of peroxidasins are well separated from other subfamilies and closely related to each other (Zamocky et al., 2008). The profile of the reconstructed phylogenetic tree suggests that vertebrate peroxidases (including MPO) evolved from a primitive peroxidasin ancestor by duplication of the peroxidase domain and loss of the residual protein part containing the leucine-rich and immunoglobulin-like motifs.
Analysis of genome databases showed the existence of two human peroxidasin genes (63% identity) located on chromosome 2p25 and 8q11, respectively. Based on highest tissue expression in heart and vascular wall (VPO1) and mainly in heart (VPO2), the peroxidasins have been designated as vascular peroxidases (VPO) (Cheng et al., 2008). The peroxidase homology region of VPO1 shows 42% identity with MPO, thus it is likely that these heme oxidoreductases carry out peroxidative reactions previously attributed exclusively to MPO in the vascular system. This hypothesis is strongly supported by critical sequence analysis based on our knowledge of structure-function relationships of mammalian peroxidases. However, so far neither structural nor detailed biochemical and biophysical data of human peroxidasins are available, yet these enzymes might play a role in defense against microbes that enter the vasculature, and also in inflammatory diseases (Cheng et al., 2008).
Aims and methods.
The cDNAs of the peroxidase domains of both human peroxidasins will be cloned, and plasmids transfected into CHO and HEK 293 cell lines. Construction of stable expression cell lines will be done in close cooperation with the Division of Mammalian Cell Culture Technology (KUNERT) based on published data performed by Cheng et al. (2008). Heme insertion into apoprotein during expression in animal cell lines by various cultivation conditions and media compositions (e.g. addition of δ-aminolevulinic acid, hemin, sodium butyrate ect.) will be tested and optimized. Heme insertion, (putative) heme to protein linkages and other post-translational modifications will be assessed by UV-Vis spectroscopy as well as liquid chromatography (LC) combined to both elemental and molecular mass spectrometry. Analysis will be carried out either on the intact protein level or on the peptide level after deglycosylation and proteolytic digestion. Heme binding will be investigated on a quantitative basis by LC coupled to inductively coupled plasma mass spectrometry (ICP-MS) that allows quantification of metal incorporation using metal-to-sulphur ratios (Hann et al. 2006).
Homology molecular models for both peroxidase domains will be performed in collaboration with OOSTENBRINK. In addition, X-ray crystallographic studies are planned in close cooperation with Ignacio FITA (Barcelona, Spain). For both recombinant oxidoreductases comprehensive spectral analyses (UV-Vis, CD- and RR-spectroscopy) of ferrous and ferric forms in the pH range 4-10 will give insights into the heme cavity architecture, spin-states, heme ligands, (putative) heme modification(s), and protein stability. Redox thermodynamic investigations will give further insights into redox mulation in peroxidasins (Battistuzzi et al., 2006).
Comprehensive steady-state kinetic analysis and determination of kinetic parameters must be performed with a set of one- and two-electron donors since corresponding data of peroxidasins are missing, and the endogeneous substrate(s) are unknown. These spectroscopic and polarographic measurements will be accompanied by time-resoved multi-mixing UV-Vis and CD-stopped-flow studies in order to analyse the kinetics of interconversion and spectroscopic features of redox intermediates (compounds I, II & III). In addition, reduction potentials of relevant short-lived redox intermediates will be determined. Functional data will be critically analysed with respect to obtained structural findings and published data of peroxidases from chordata peroxidases (subfamily 1).
Battistuzzi, G., Bellei, M., Zederbauer, M., Furtmüller, P. G., Sola, M., Obinger, C. (2006) Redox thermodynamics of the Fe(III)/Fe(II) couple of human myeloperoxidase in its high-spin and low-spin forms. Biochemistry 45, 12750-12755
Cheng, G., Salerno, J. C., Cao, Z., Pagano, P. J., Lambeth, J. D. (2008) Identification and characterization of VPO1, a new animal heme-containing peroxidase. Free Rad. Biol. Med. 45, 1682-1694
Hann, S., Obinger, C., Stingeder, G., Paumann, M., Furtmüller, P. G., Koellensperger, G. (2006) Studying metal integration in native and recombinant copper proteins by hyphenated ICP-DRC-MS and ESI-TOF-MS capabilities and limitations of the complementary techniques. J. Anal. At. Spectrom. 21, 1224-1231
Nelson, R. E., Fessler, L. I., Tkagi, Y., Blumberg, B., Keene, D. R., Olson, P. F., Parker, C. G., Fessler, J. H. (1994) Peroxidasin: a novel enzyme-matrix protein of Drosophila development. EMBO J. 13, 3438-3447
Zamocky, M., Jakopitsch, C., Furtmüller, P. G., Dunand, C., Obinger, C. (2008) The peroxidase-cyclooxygenase superfamily. Reconstructed evolution of critical enzymes of the innate immune defense system. Proteins 71, 589-605
