Biocatalysis of chlorite dismutation – functional and structural studies on new peculiar heme enzymes
SUPERVISOR: CHRISTIAN OBINGER
Project assigned to: STEFAN HOFBAUER
Background.
The heme enzyme chlorite dismutase (Cld) was discovered in chlorate- and perchlorate-reducing (proteo)bacteria (van Ginkel et al., 1996), which are facultative anaerobes using chlorate or perchlorate as alternative electron acceptors in the absence of oxygen. The reduction of (per)chlorate yields chlorite, which is a strong oxidant and has cell-damaging effects. At this point Cld comes into play: this metalloenzyme degrades chlorite to chloride and dioxygen and thus protects these organisms from the accumulation of harmful chlorite. From a biochemical point of view Cld is an exciting oxidoreductase since – besides photosystem II – so far Cld is the only known enzyme that is able to catalyze the formation of a covalent O-O bond, which leads to the synthesis of dioxygen. However, the catalytic mechanism of chlorite degradation and O2 formation by Cld is not fully understood yet (Lee et al., 2008). Cld is also of high practical relevance for the bioremediation of the anthropogenic pollutants (per)chlorate and chlorite. These toxic compounds are released into the environment due to industrial processes and the application of fertilizers, disinfectants and bleaching agents.
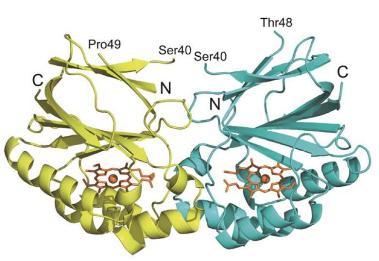
Recently, highly active Cld was also identified and found to be expressed in vivo in the nitrite-oxidizing bacterium “Candidatus Nitrospira defluvii”, a member of the bacterial phylum Nitrospira (Maixner et al., 2008), an aerobic and chemolithoautotrophic organism that gains energy from the oxidation of nitrite to nitrate. This raises new and fundamental questions regarding the evolution of Cld and its biological role. Furthermore, recent genome analysis revealed a relatively large superfamily of Cld-like proteins, which extents over a surprising number of bacterial and also archaeal phyla. The known bona fide Clds from chlorate- and perchlorate-reducing bacteria constitute only a minor group within this superfamily. The enzymatic activities and functional roles of the vast majority of the Cld-like proteins are completely unknown. Moreover, there seems to be an unexpected broad structural (oligomerization, subunit size, substrate accessibility) and functional diversity (enzyme kinetics) of these metalloproteins that is not understood yet (Kostan et al., 2010; Mlynek et al., 2010). Its understanding is a precondition for application of these biocatalysts in biotechnology.
Aims and methods.
Heterologous recombinant expression in E. coli of homopentameric Cld from “Candidatus Nitrospira defluvii” and homodimeric Cld from “Nitrobacter winogradskyi”. Based on recently solved X-ray structures (Kostan et al., 2010; Mlynek et al., 2010) site-directed mutagenesis of distal and proximal amino acids will be performed. Emphasis will be placed on amino acids involved in substrate binding and oxidation, redox regulation and internal electron transfer.
Mutant proteins will be characterized by a broad set of biochemical/physical methods including (i) X-ray crystallography (in close cooperation with Kristina DJINOVIC-CARUGO from the Department for Structural and Computational Biology, Max F. Perutz Laboratories, University of Vienna), (ii) detailed spectral analysis (UV-Vis and resonance Raman) of proteins in different redox- and spin-states, (iii) time-resoved multi-mixing UV-Vis and CD-stopped-flow studies in order to analyse the kinetics of interconversion and spectroscopic features of relevant redox intermediates, (iV) spectroelectrochemical studies (in cooperation with Gianantonio BATTISTUZZI from the Department of Chemistry, University of Modena and Reggio Emilia, Italy). In addition it is planned to analyze the formation and migration of protein radical sites by freeze-quence electron paramagnetic resonance spectroscopy.
These methods will provide valuable insight into the mechanism of chlorite degradation and proposed pathways will be analyzed by classical and ab initio molecular dynamics simulations (in cooperation with OOSTENBRINK). Obtained data will provide an excellent basis for further engineering projects (e.g. stability engineering) on these peculiar metalloenzymes for usage in bioremediation .
Lee, A.Q., Streit, B.R., Zdilla, M.J., Abu-Omar, M.M., DuBois, J.L. (2008) Mechanism of and exquisite selectivity for O-O bond formation by the heme-dependent chlorite dismutase. Proc. Natl. Acad. Sci. USA 105, 15654-15659
Kostan, J., Sjöblom, B., Maixner, F., Mlynek, G., Furtmüller, P.G., Obinger, C., Wagner, M., Daims, H., Djinovic-Carugo, K. (2010) Structural and functional characterisation of the chlorite dismutase from the nitrite-oxidizing bacterium “Candidatus Nitrospira defluvii”: identification of a catalytically important amino acid residue. J. Struct. Biol., in press
Maixner, F., Wagner, M., Lucker, S., Pelletier, E., Schmitz-Esser, S., Hace, K., Spieck, E., Konrat, R., Le Paslier, D., Daims, H. (2008) Environmental genomics reveals a functional chlorite dismutase in the nitrite-oxidizing bacterium “Candidatus Nitrospira defluvii”. Environ. Microbiol. 10, 3043-3056
Mlynek, G., Sjöblom, B., Kostan, J., Füreder, S., Maixner, F., Furtmüller, P.G., Obinger, C., Wagner, M., Daims, H., Djinovic-Carugo, K. (2010) Unexpected diversity of chlorite dismutases: a catalytically efficient dimeric enzyme from Nitrobacter winogradskyi. Submitted for publication
Van Ginkel, C.G., Rikken, G.B., Kroon, A.G., Kengen, S.W. (1996) Purification and characterization of chlorite dismutase: a novel oxygen-generating enzyme. Arch. Microbiol. 166, 321-326
